healthyverse_tsa
Time Series Analysis, Modeling and Forecasting of the Healthyverse Packages
Steven P. Sanderson II, MPH - Date: 2026-04-29
Introduction
This analysis follows a Nested Modeltime Workflow from modeltime
along with using the NNS package. I use this to monitor the
downloads of all of my packages:
Get Data
glimpse(downloads_tbl)
Rows: 176,636
Columns: 11
$ date <date> 2020-11-23, 2020-11-23, 2020-11-23, 2020-11-23, 2020-11-23,…
$ time <Period> 15H 36M 55S, 11H 26M 39S, 23H 34M 44S, 18H 39M 32S, 9H 0M…
$ date_time <dttm> 2020-11-23 15:36:55, 2020-11-23 11:26:39, 2020-11-23 23:34:…
$ size <int> 4858294, 4858294, 4858301, 4858295, 361, 4863722, 4864794, 4…
$ r_version <chr> NA, "4.0.3", "3.5.3", "3.5.2", NA, NA, NA, NA, NA, NA, NA, N…
$ r_arch <chr> NA, "x86_64", "x86_64", "x86_64", NA, NA, NA, NA, NA, NA, NA…
$ r_os <chr> NA, "mingw32", "mingw32", "linux-gnu", NA, NA, NA, NA, NA, N…
$ package <chr> "healthyR.data", "healthyR.data", "healthyR.data", "healthyR…
$ version <chr> "1.0.0", "1.0.0", "1.0.0", "1.0.0", "1.0.0", "1.0.0", "1.0.0…
$ country <chr> "US", "US", "US", "GB", "US", "US", "DE", "HK", "JP", "US", …
$ ip_id <int> 2069, 2804, 78827, 27595, 90474, 90474, 42435, 74, 7655, 638…
The last day in the data set is 2026-04-27 23:18:28, the file was birthed on: 2025-10-31 10:47:59.603742, and at report knit time is 4280.51 hours old. Happy analyzing!
Now that we have our data lets take a look at it using the skimr
package.
skim(downloads_tbl)
| Name | downloads_tbl |
| Number of rows | 176636 |
| Number of columns | 11 |
| _______________________ | |
| Column type frequency: | |
| character | 6 |
| Date | 1 |
| numeric | 2 |
| POSIXct | 1 |
| Timespan | 1 |
| ________________________ | |
| Group variables | None |
Data summary
Variable type: character
| skim_variable | n_missing | complete_rate | min | max | empty | n_unique | whitespace |
|---|---|---|---|---|---|---|---|
| r_version | 131370 | 0.26 | 5 | 7 | 0 | 51 | 0 |
| r_arch | 131370 | 0.26 | 1 | 7 | 0 | 6 | 0 |
| r_os | 131370 | 0.26 | 7 | 19 | 0 | 27 | 0 |
| package | 0 | 1.00 | 7 | 13 | 0 | 8 | 0 |
| version | 0 | 1.00 | 5 | 17 | 0 | 63 | 0 |
| country | 16475 | 0.91 | 2 | 2 | 0 | 167 | 0 |
Variable type: Date
| skim_variable | n_missing | complete_rate | min | max | median | n_unique |
|---|---|---|---|---|---|---|
| date | 0 | 1 | 2020-11-23 | 2026-04-27 | 2024-01-19 | 1975 |
Variable type: numeric
| skim_variable | n_missing | complete_rate | mean | sd | p0 | p25 | p50 | p75 | p100 | hist |
|---|---|---|---|---|---|---|---|---|---|---|
| size | 0 | 1 | 1130037.06 | 1478562.36 | 355 | 43636 | 325190 | 2334859 | 5677952 | ▇▁▂▁▁ |
| ip_id | 0 | 1 | 11564.39 | 23290.22 | 1 | 184 | 2732 | 11769 | 299146 | ▇▁▁▁▁ |
Variable type: POSIXct
| skim_variable | n_missing | complete_rate | min | max | median | n_unique |
|---|---|---|---|---|---|---|
| date_time | 0 | 1 | 2020-11-23 09:00:41 | 2026-04-27 23:18:28 | 2024-01-19 12:44:45 | 112547 |
Variable type: Timespan
| skim_variable | n_missing | complete_rate | min | max | median | n_unique |
|---|---|---|---|---|---|---|
| time | 0 | 1 | 0 | 59 | 20.5 | 60 |
We can see that the following columns are missing a lot of data and for
us are most likely not useful anyways, so we will drop them
c(r_version, r_arch, r_os)
Plots
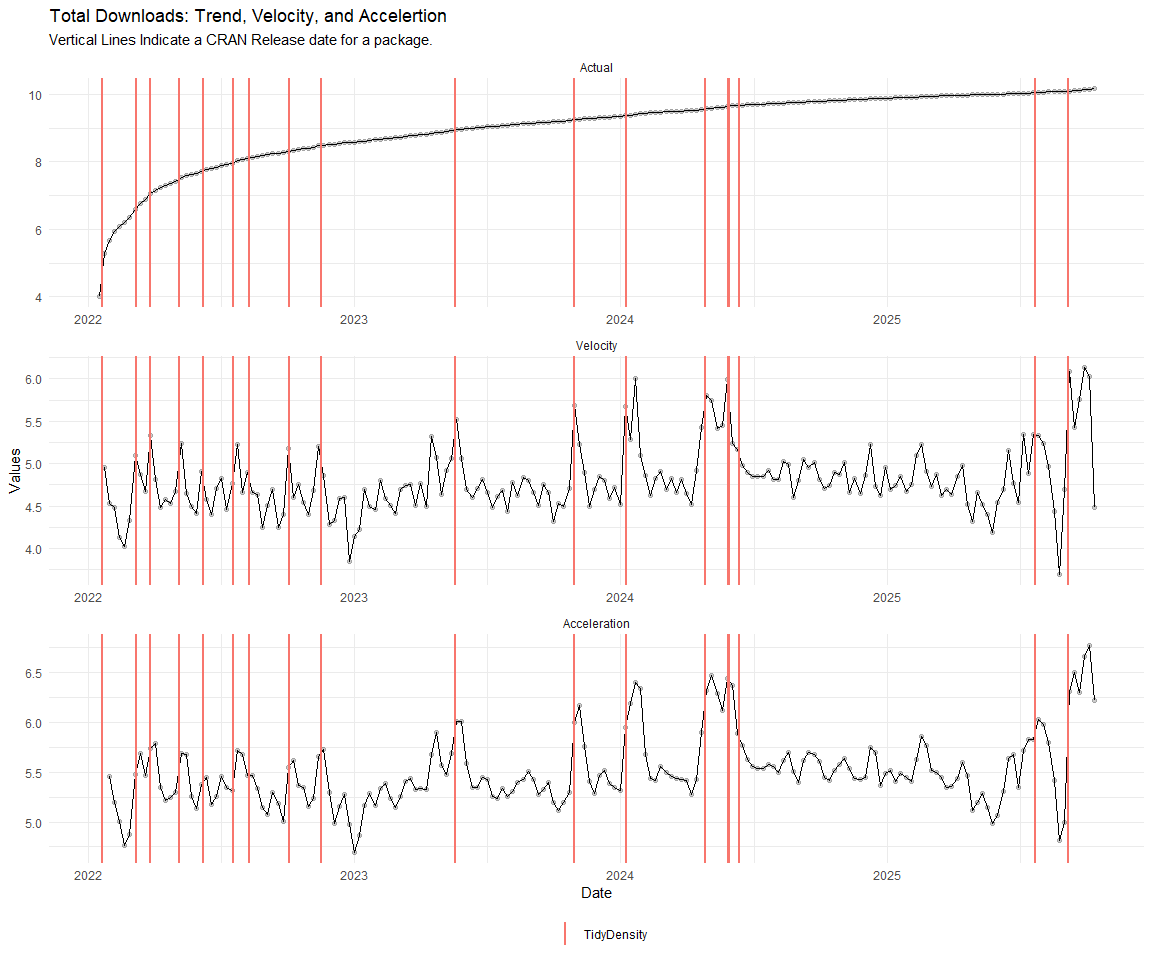
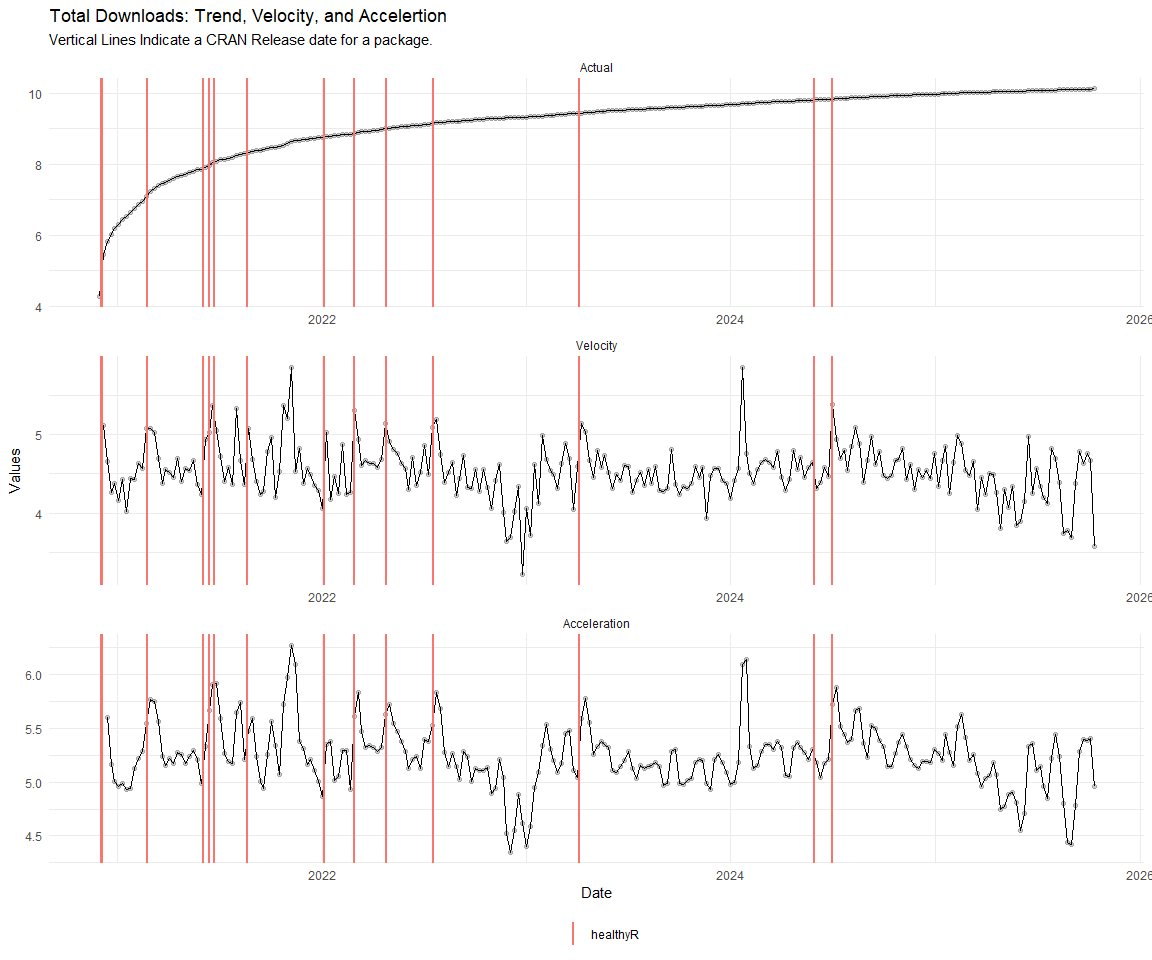
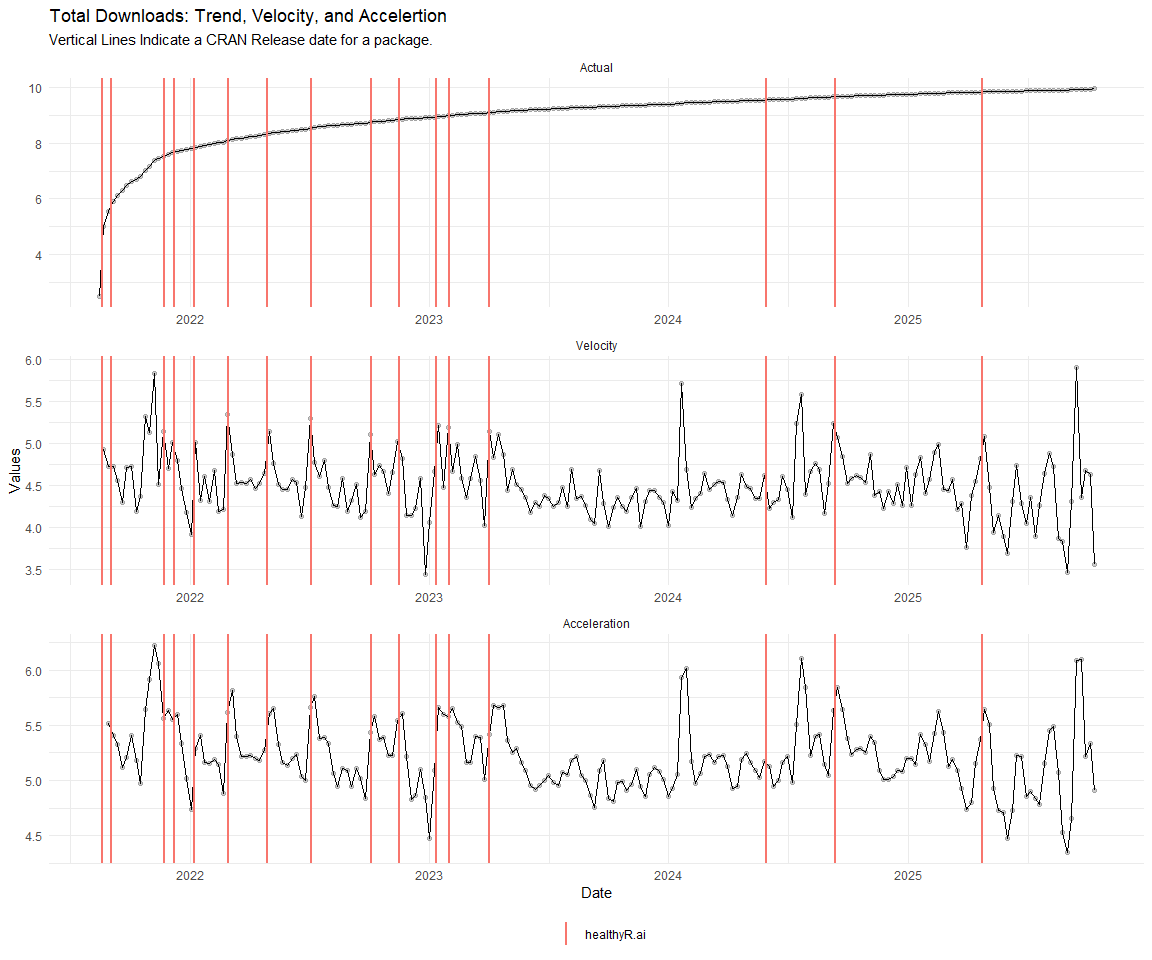
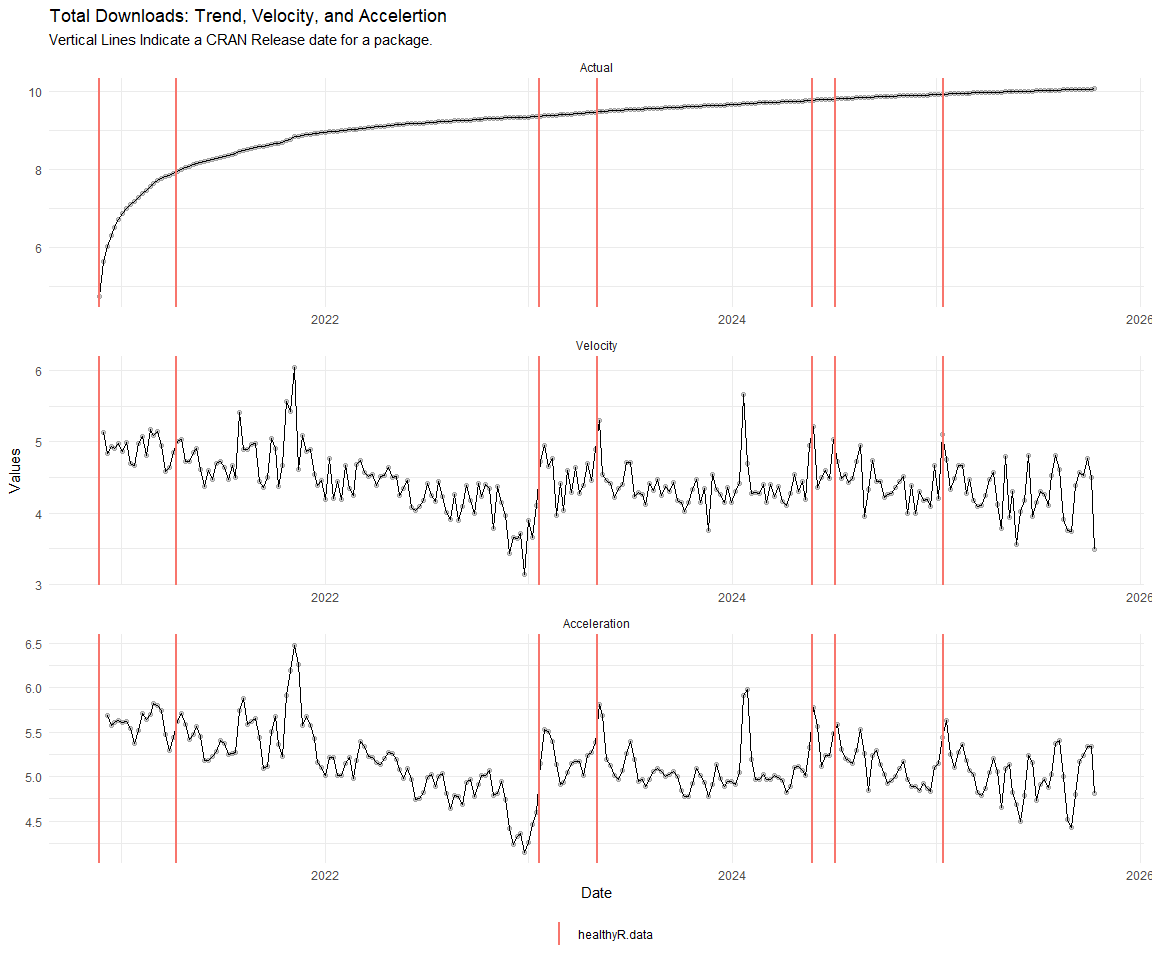
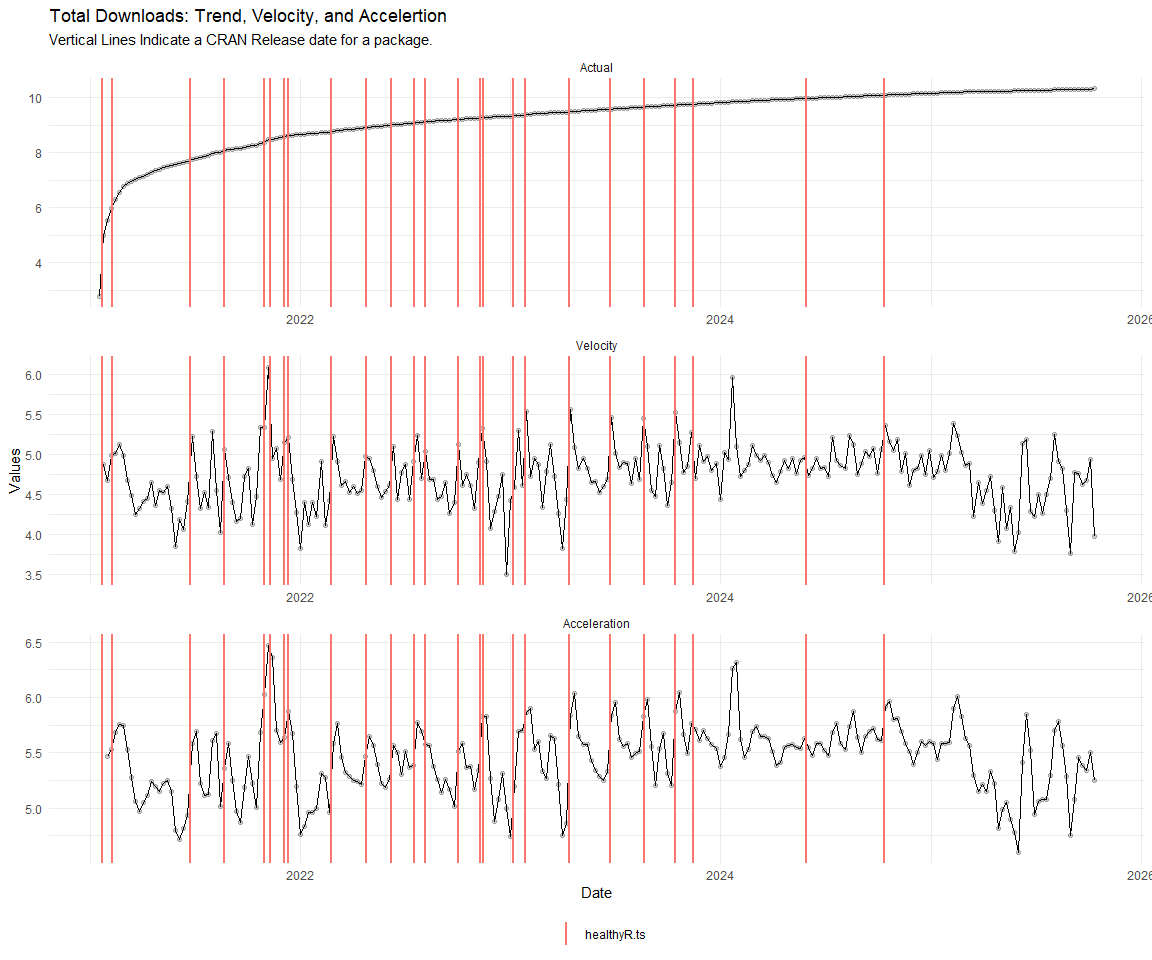
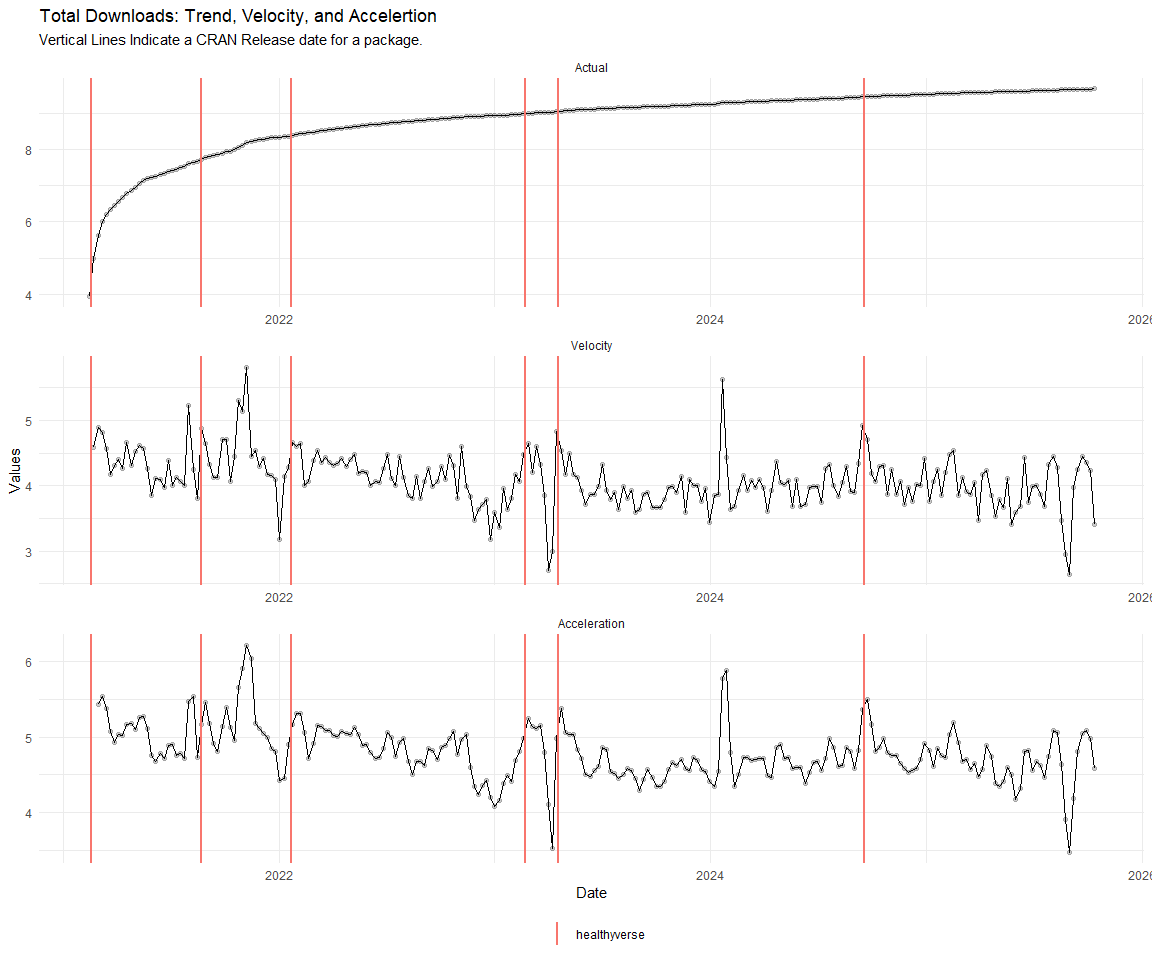
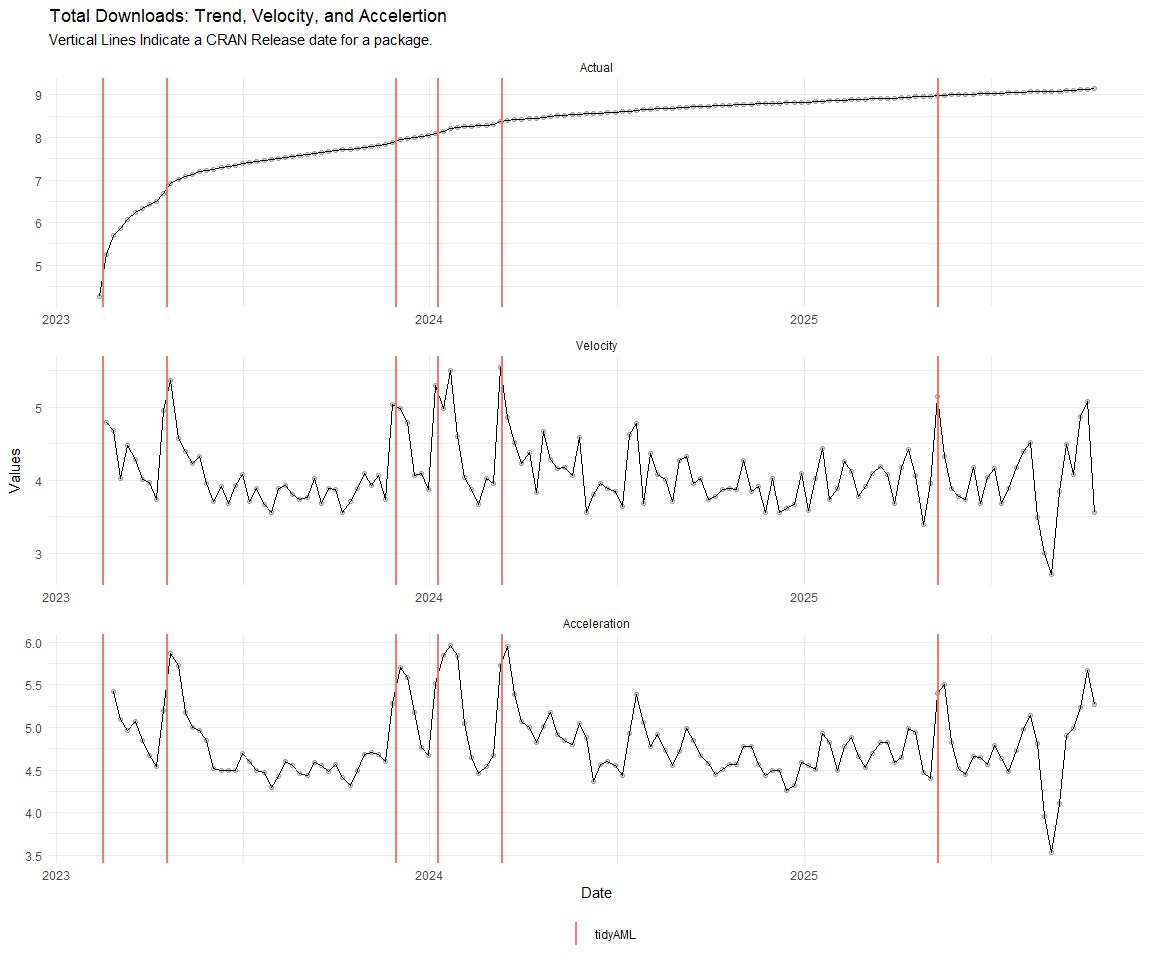
Now lets take a look at a time-series plot of the total daily downloads by package. We will use a log scale and place a vertical line at each version release for each package.
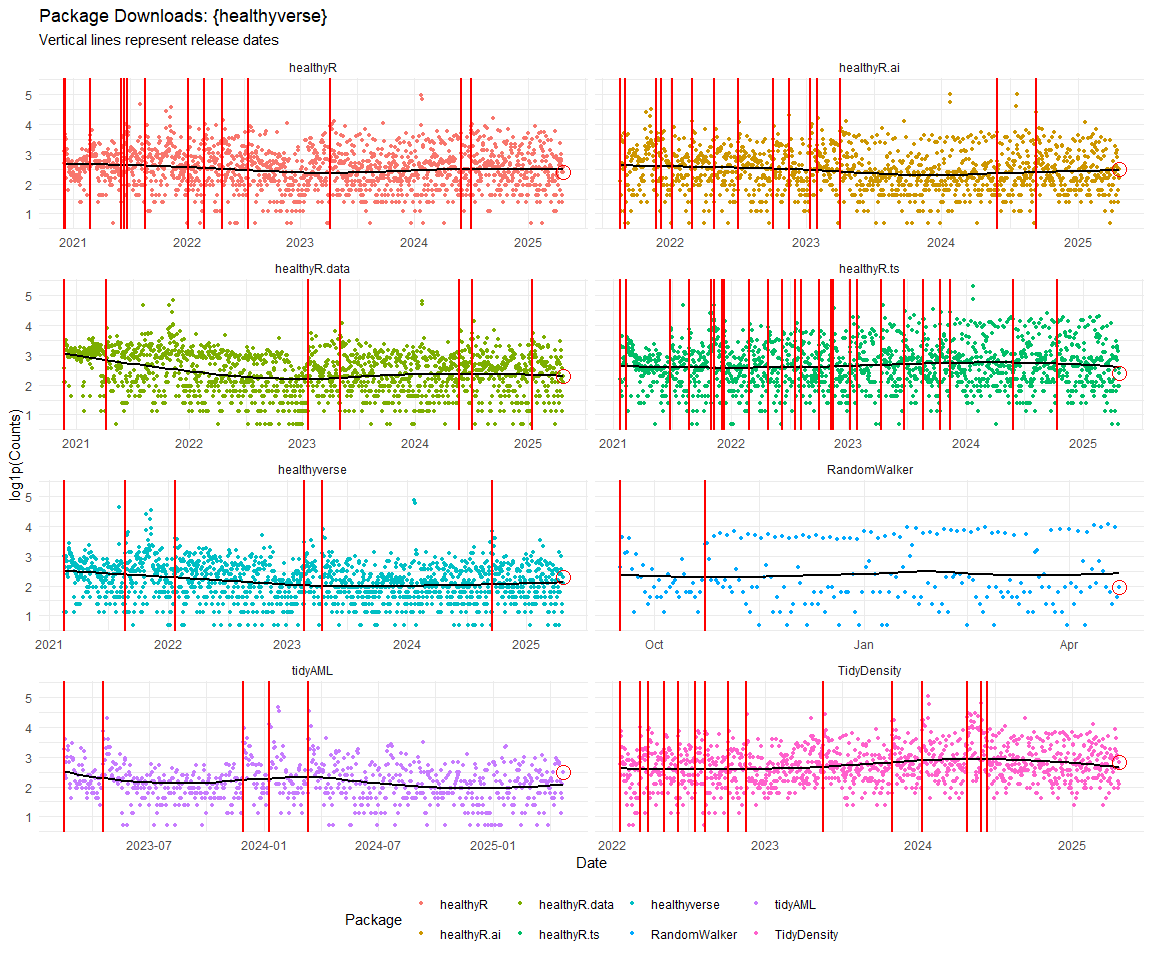
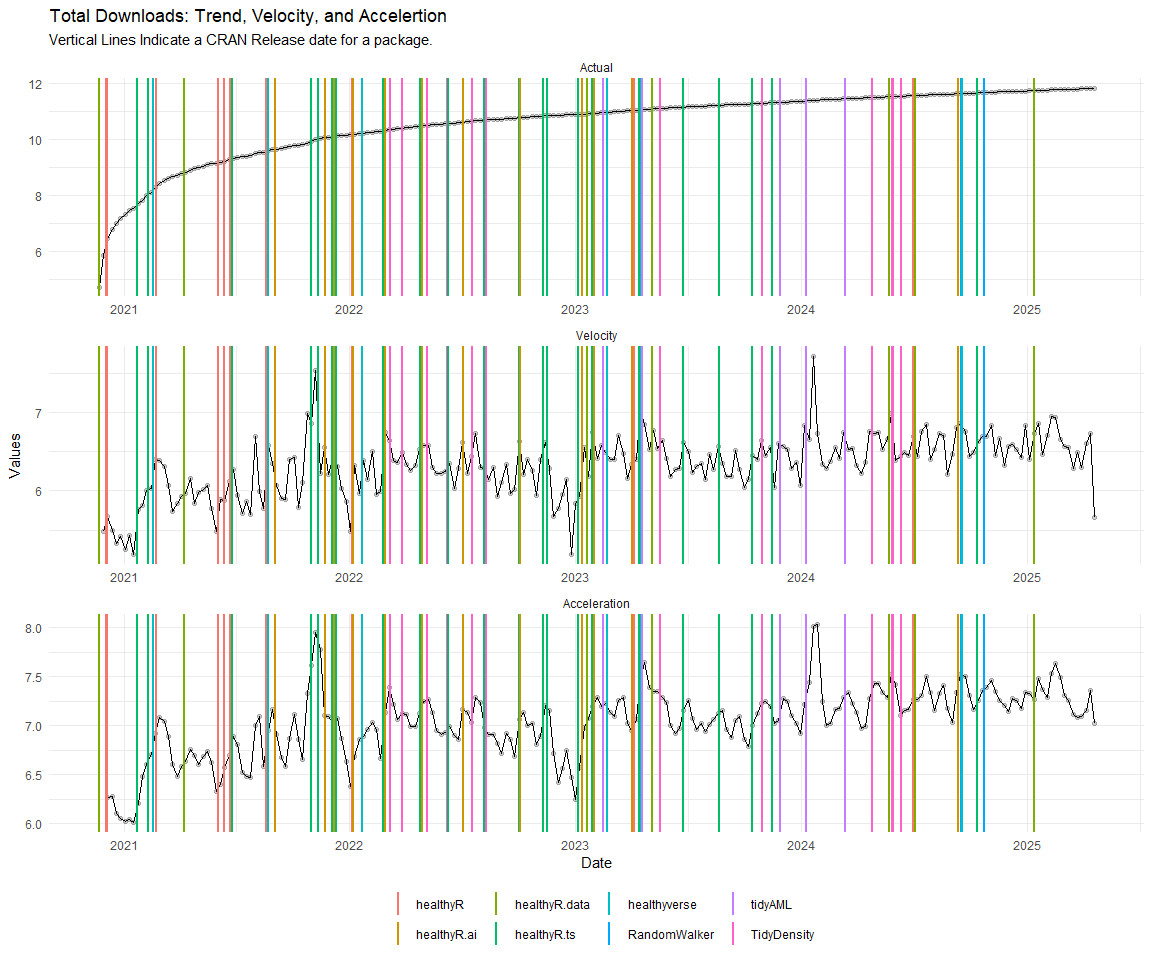
[[1]]
[[2]]
[[3]]
[[4]]
[[5]]
[[6]]
[[7]]
[[8]]
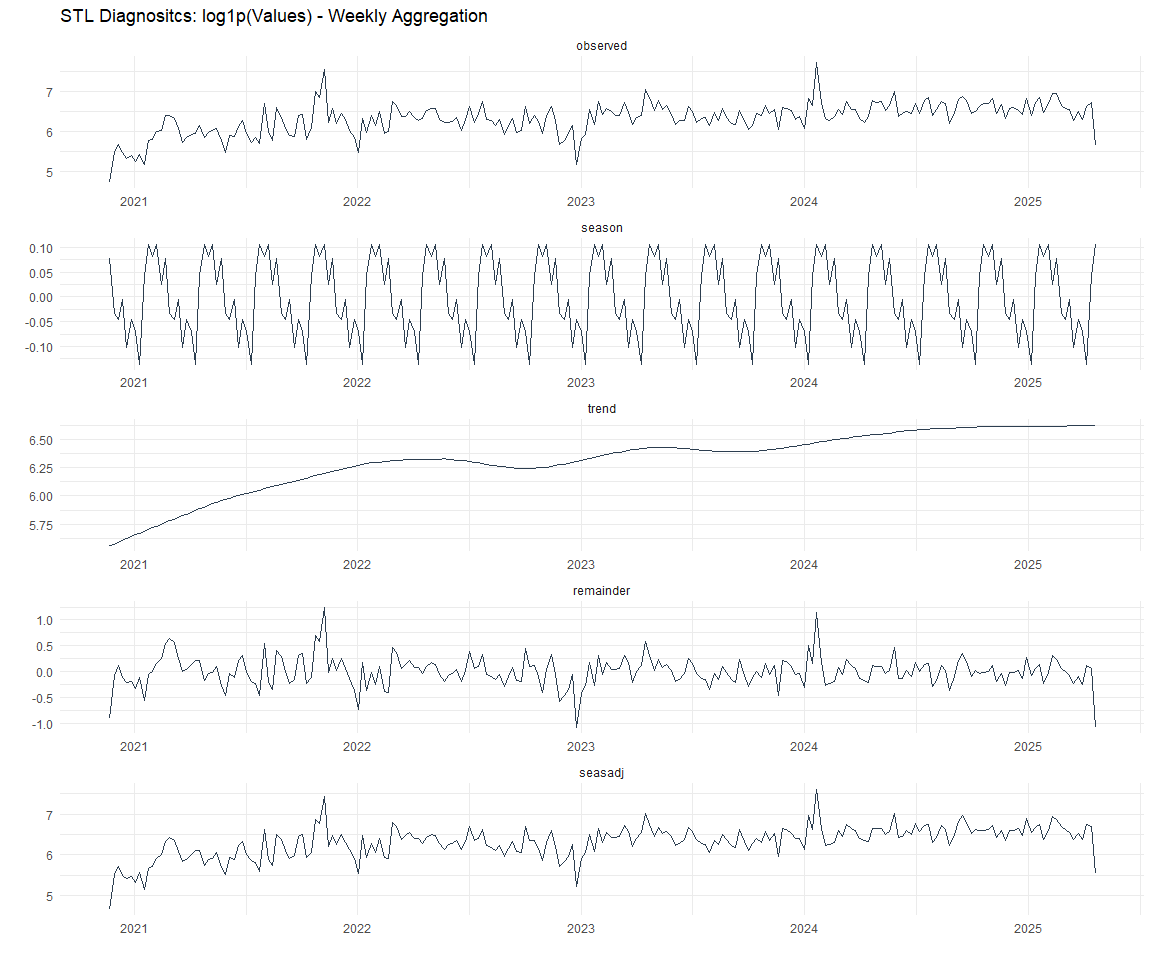
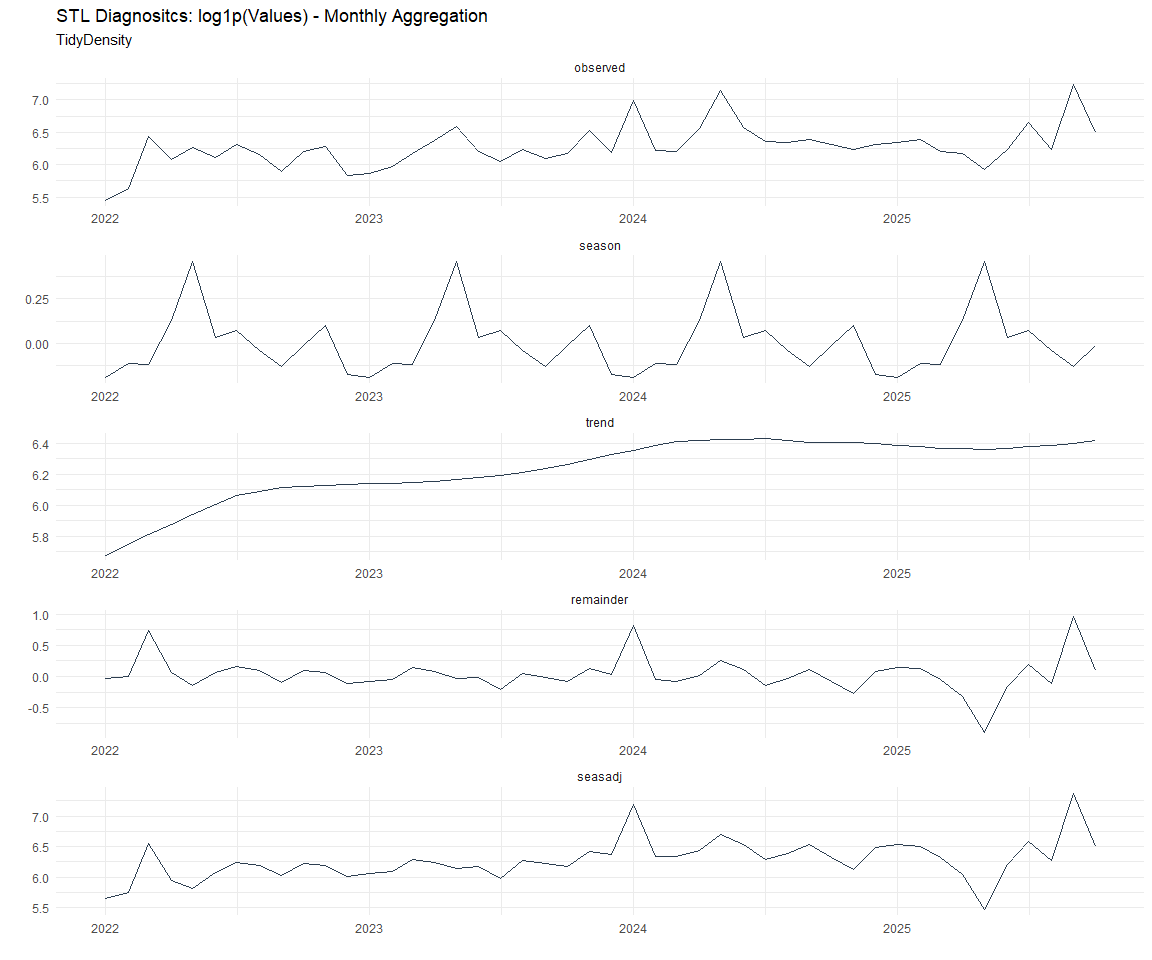
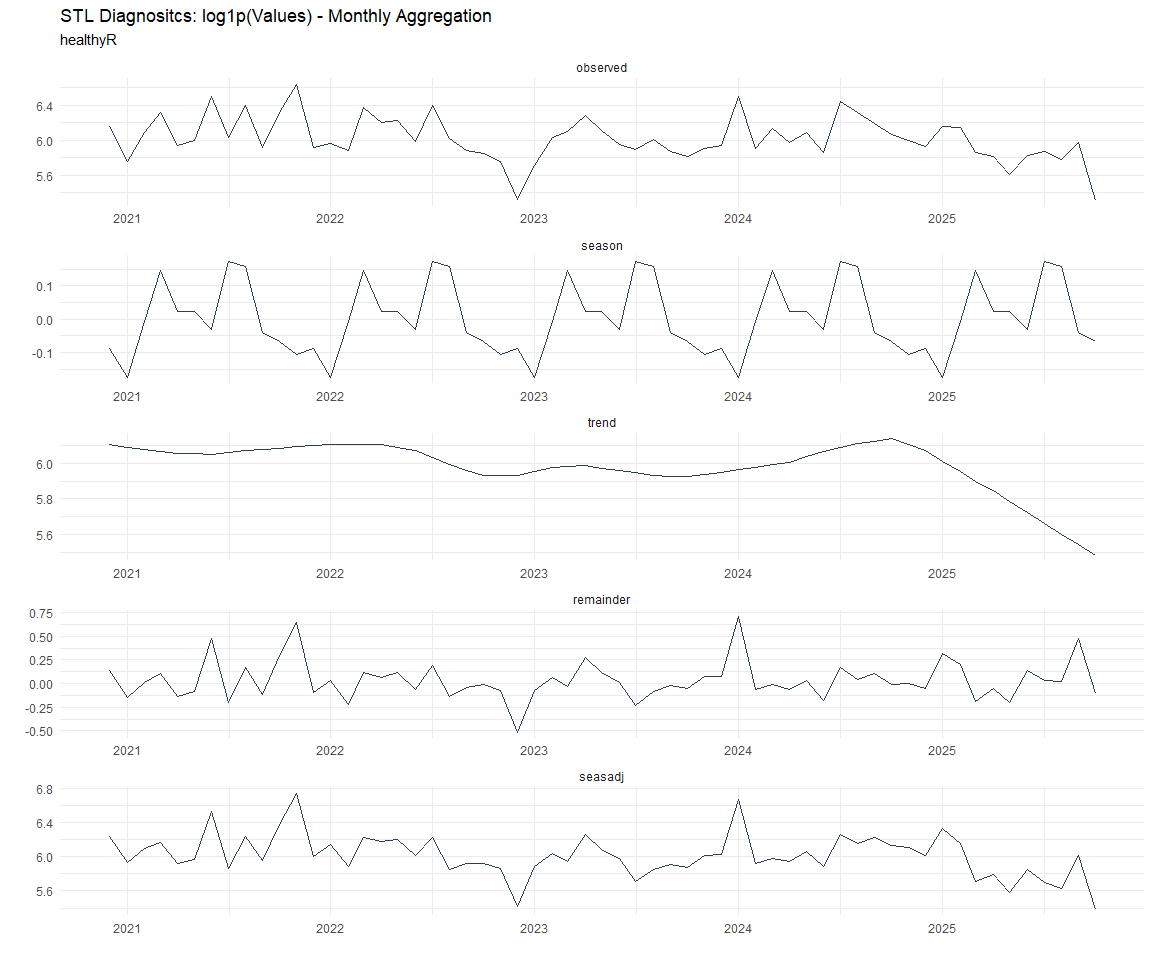
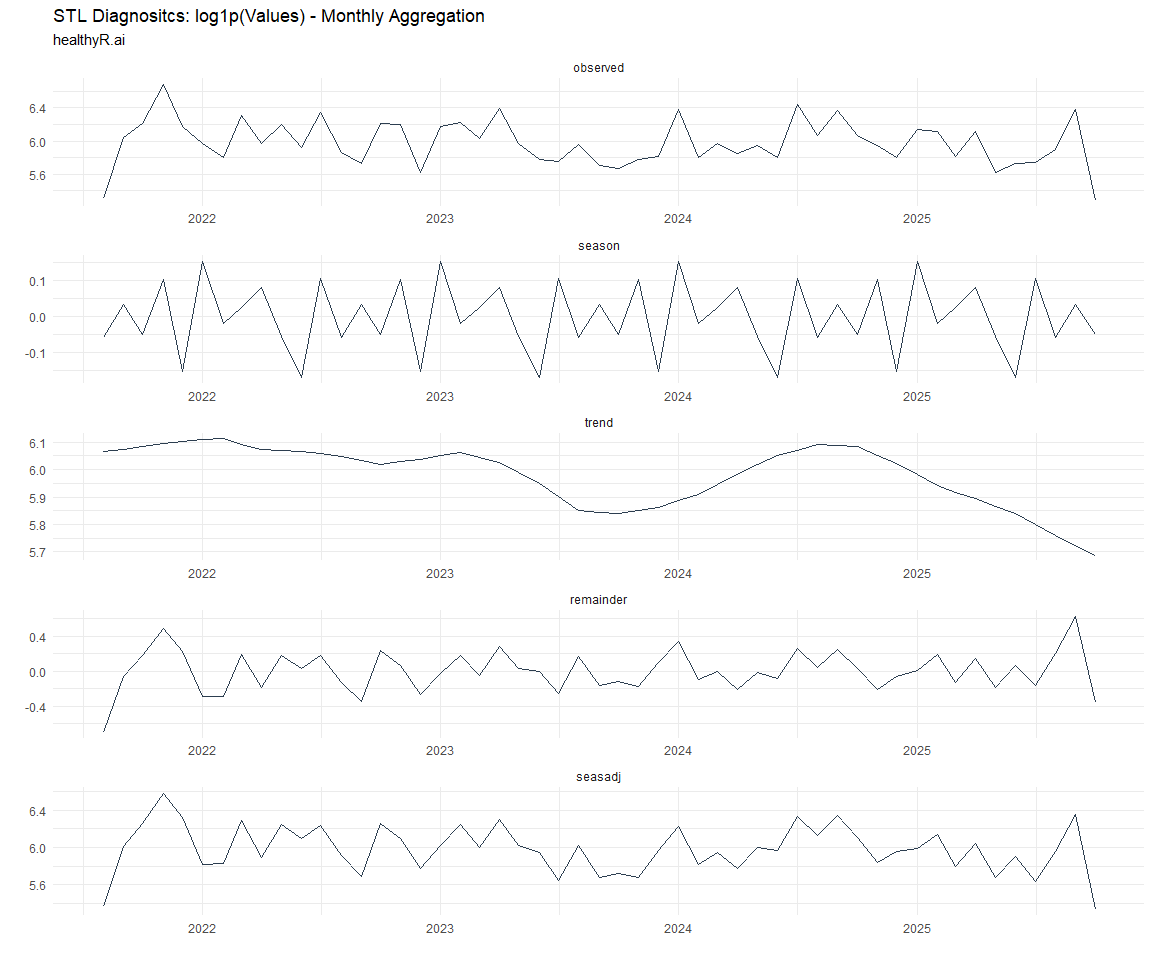
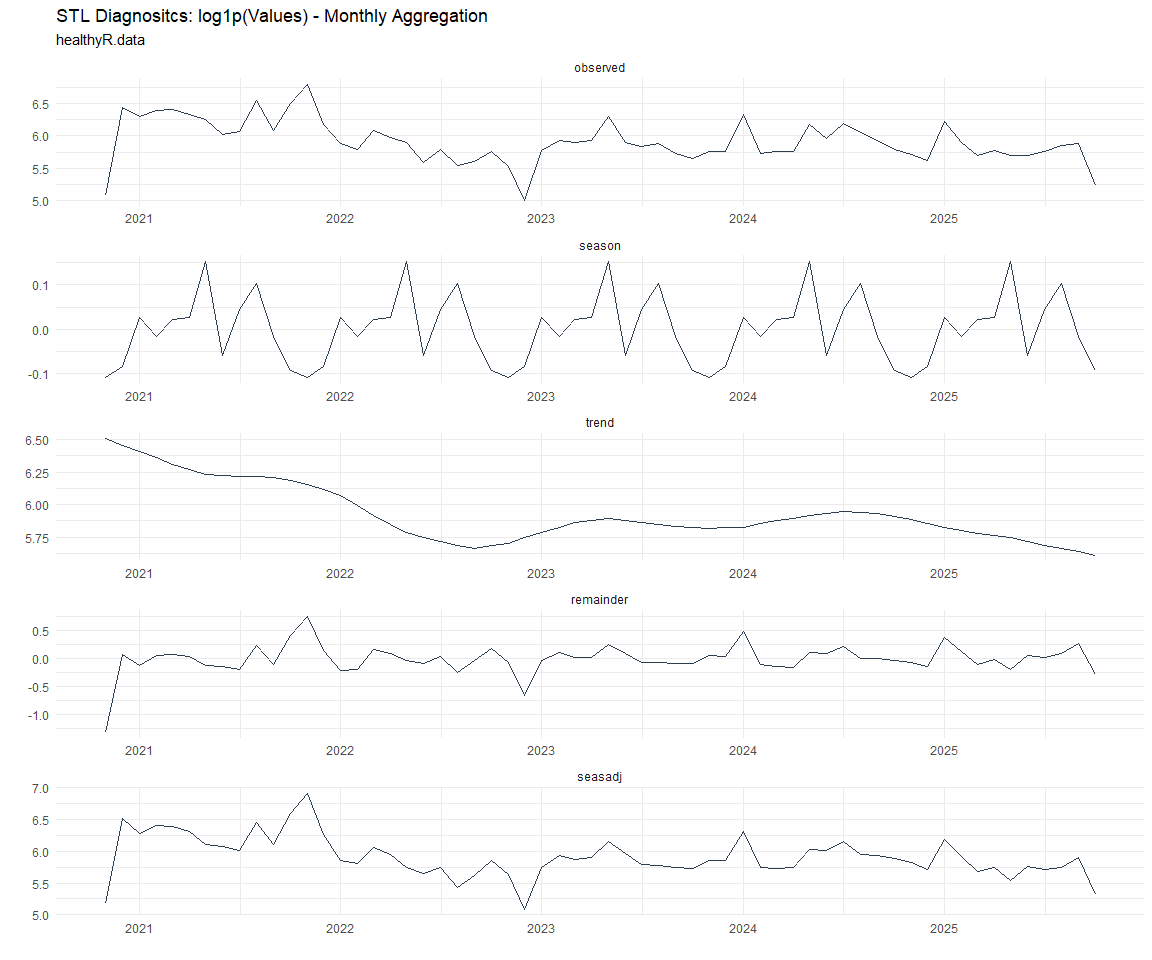
Now lets take a look at some time series decomposition graphs.
[[1]]
[[2]]
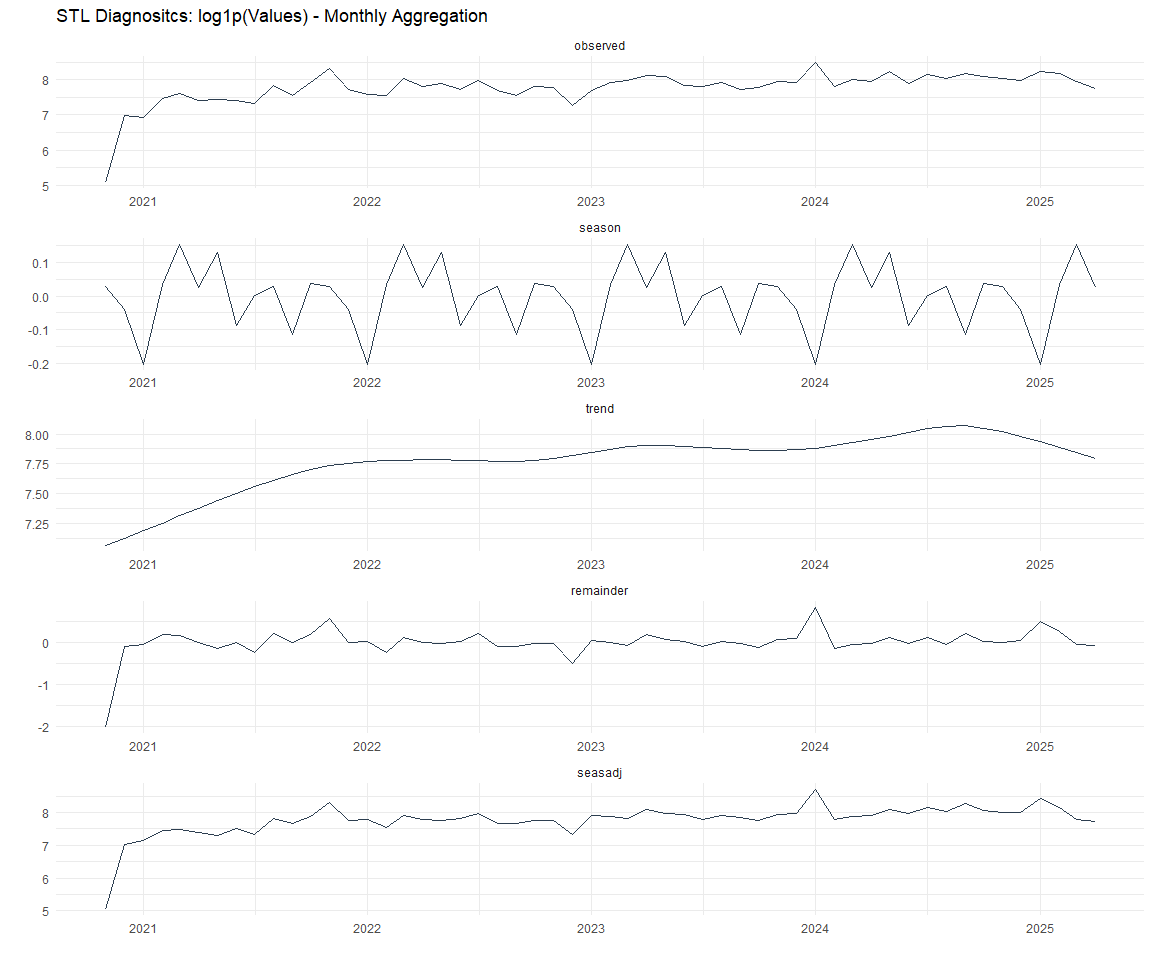
[[3]]
[[4]]
[[5]]
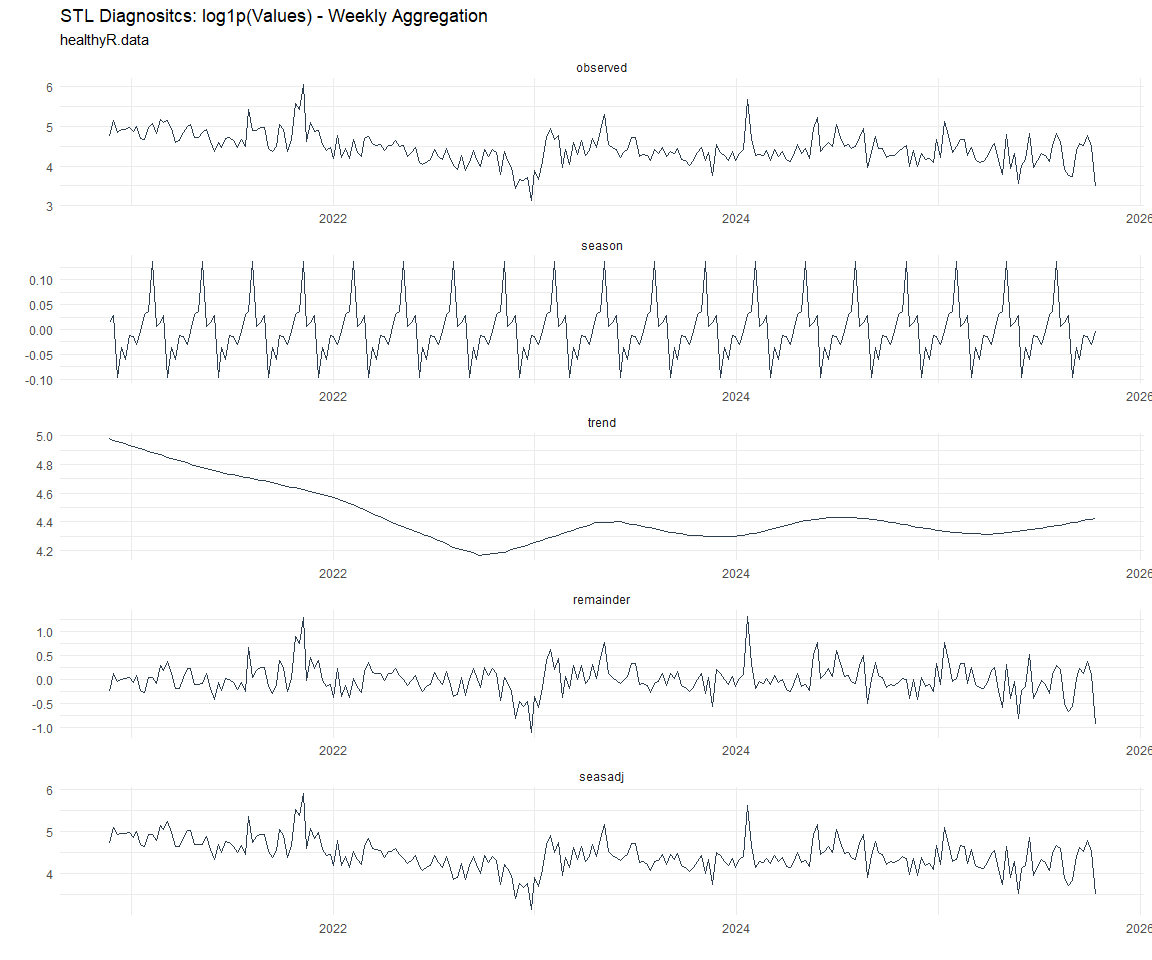
[[6]]
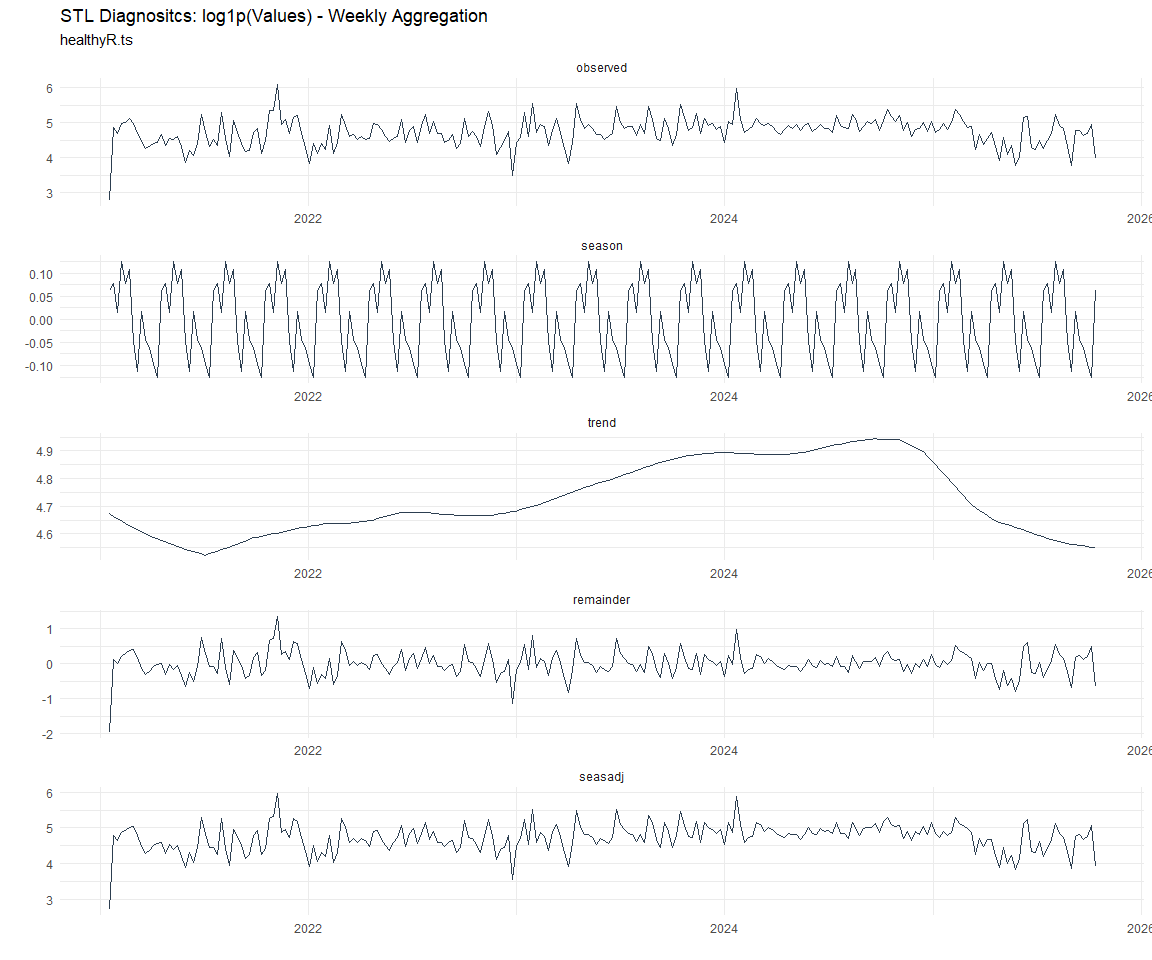
[[7]]
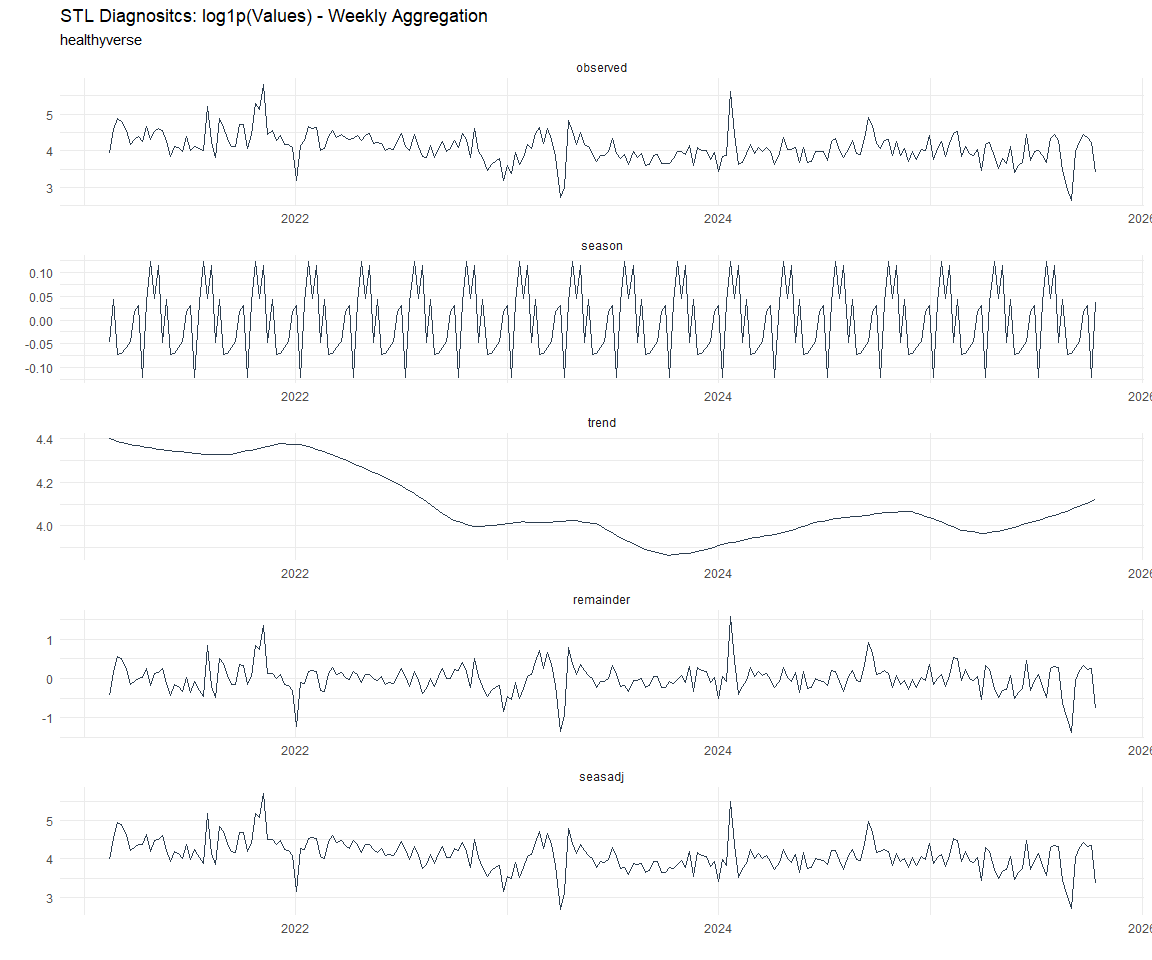
[[8]]
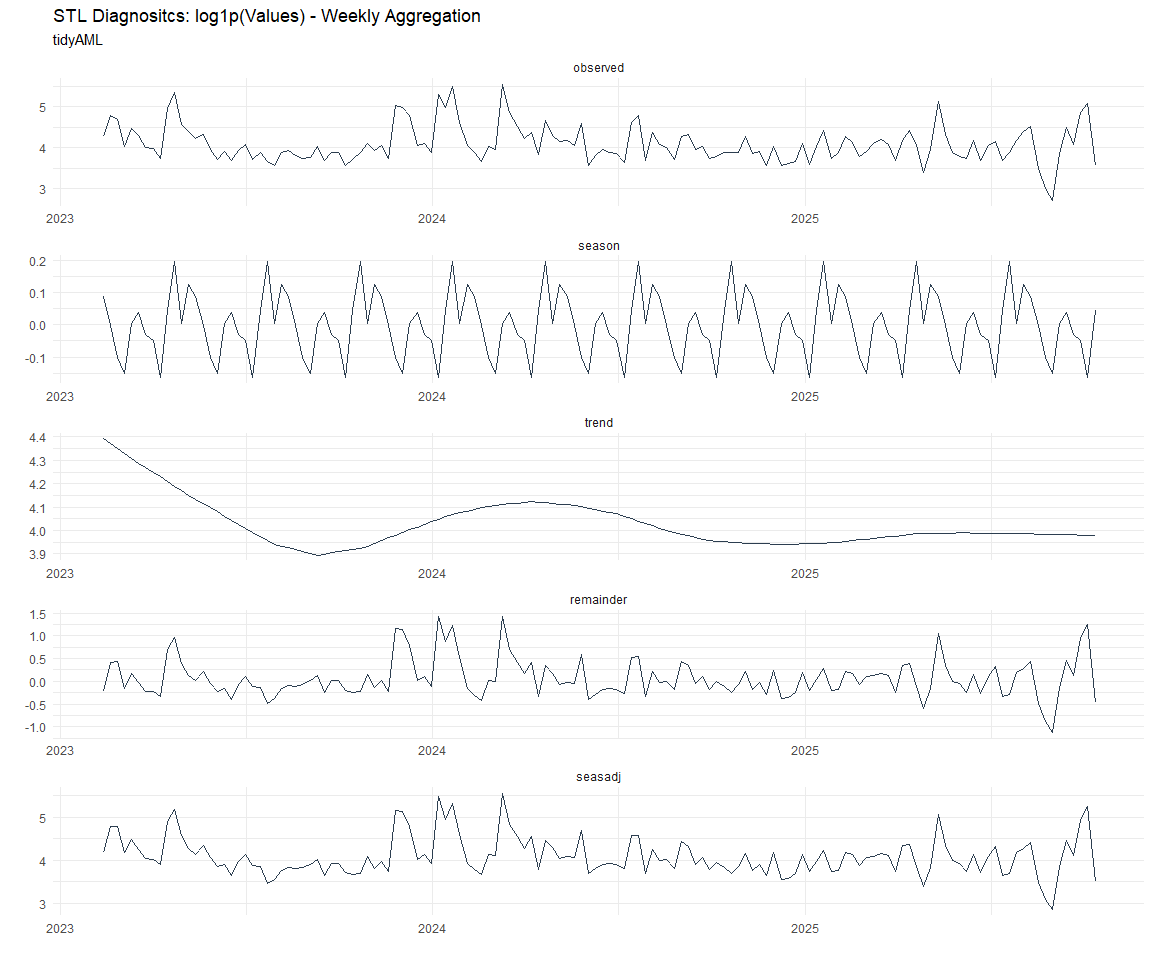
[[1]]
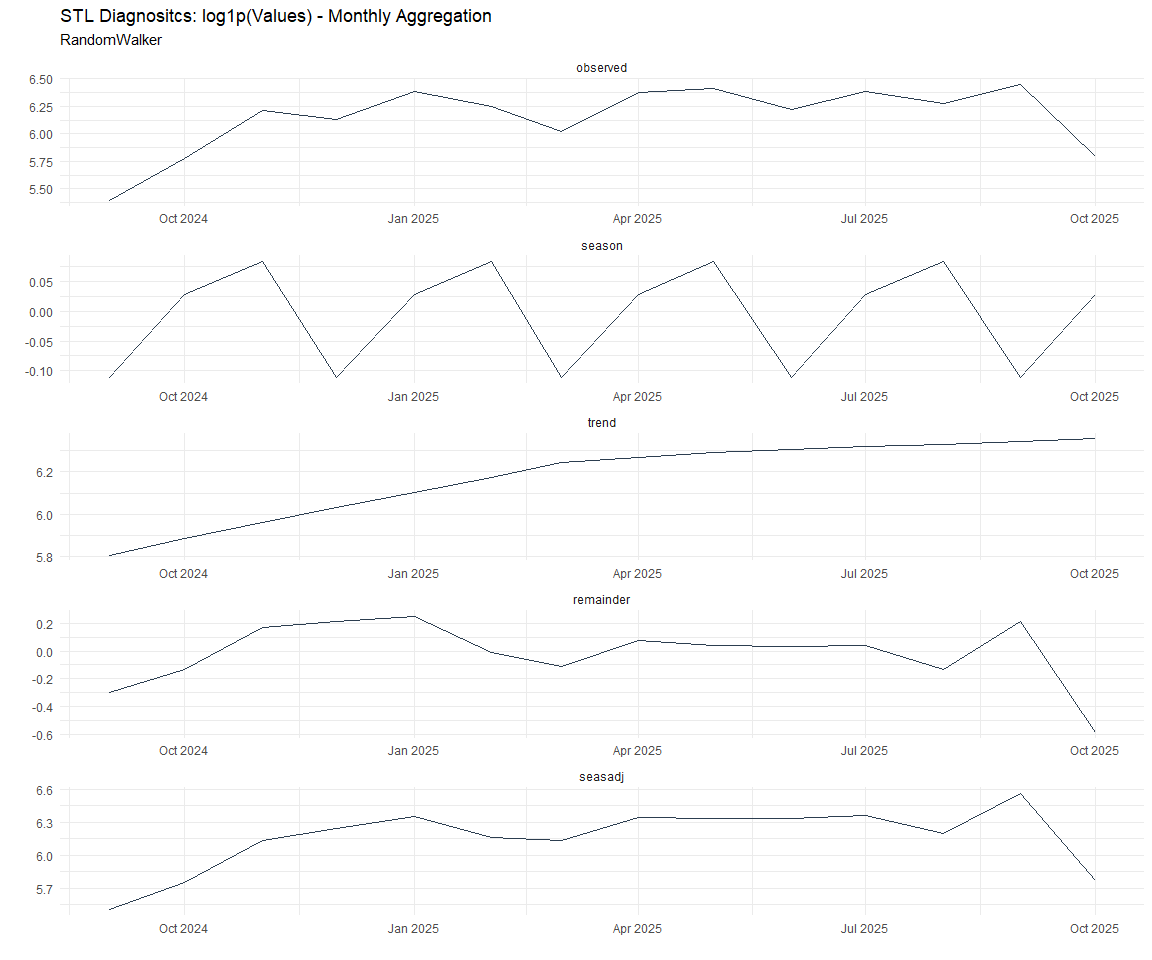
[[2]]
[[3]]
[[4]]
[[5]]
[[6]]
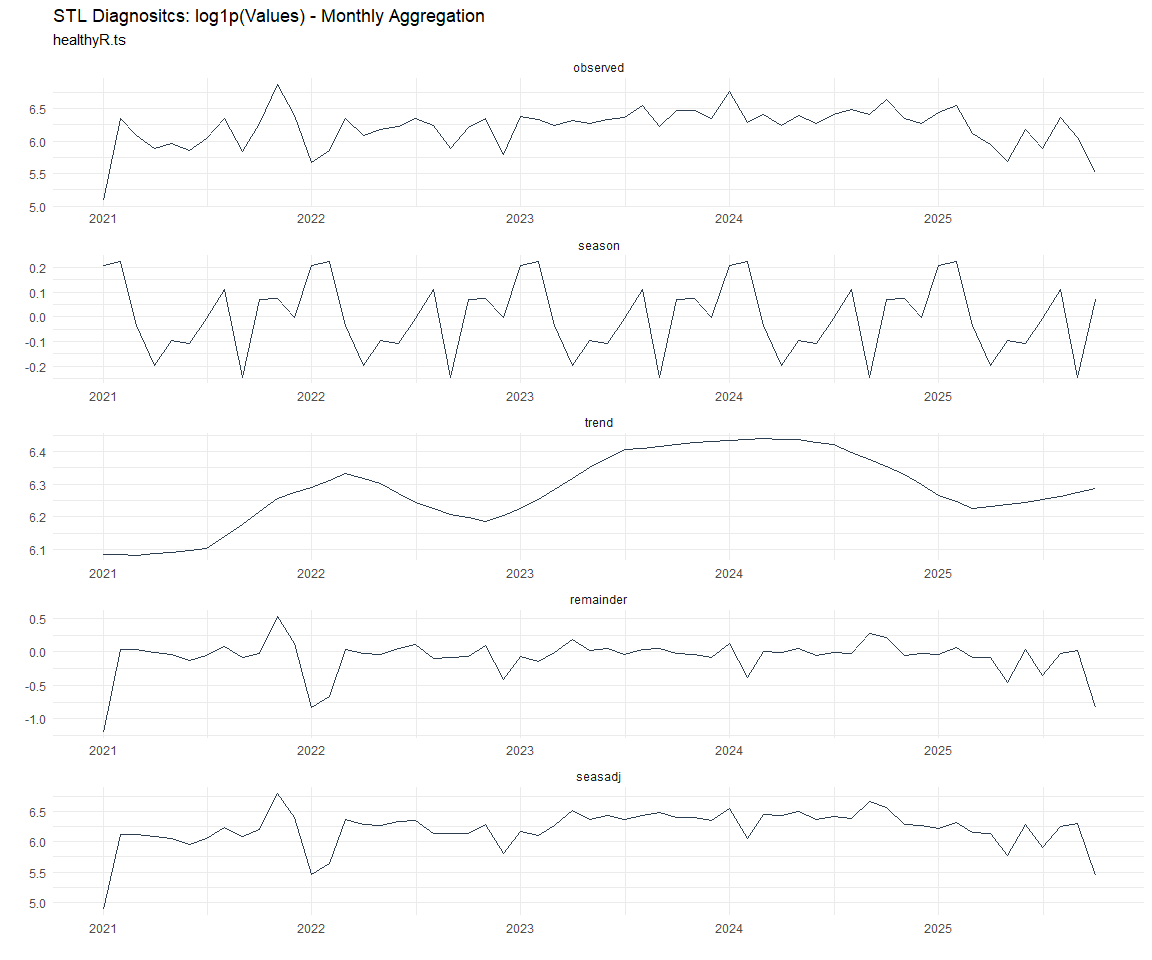
[[7]]
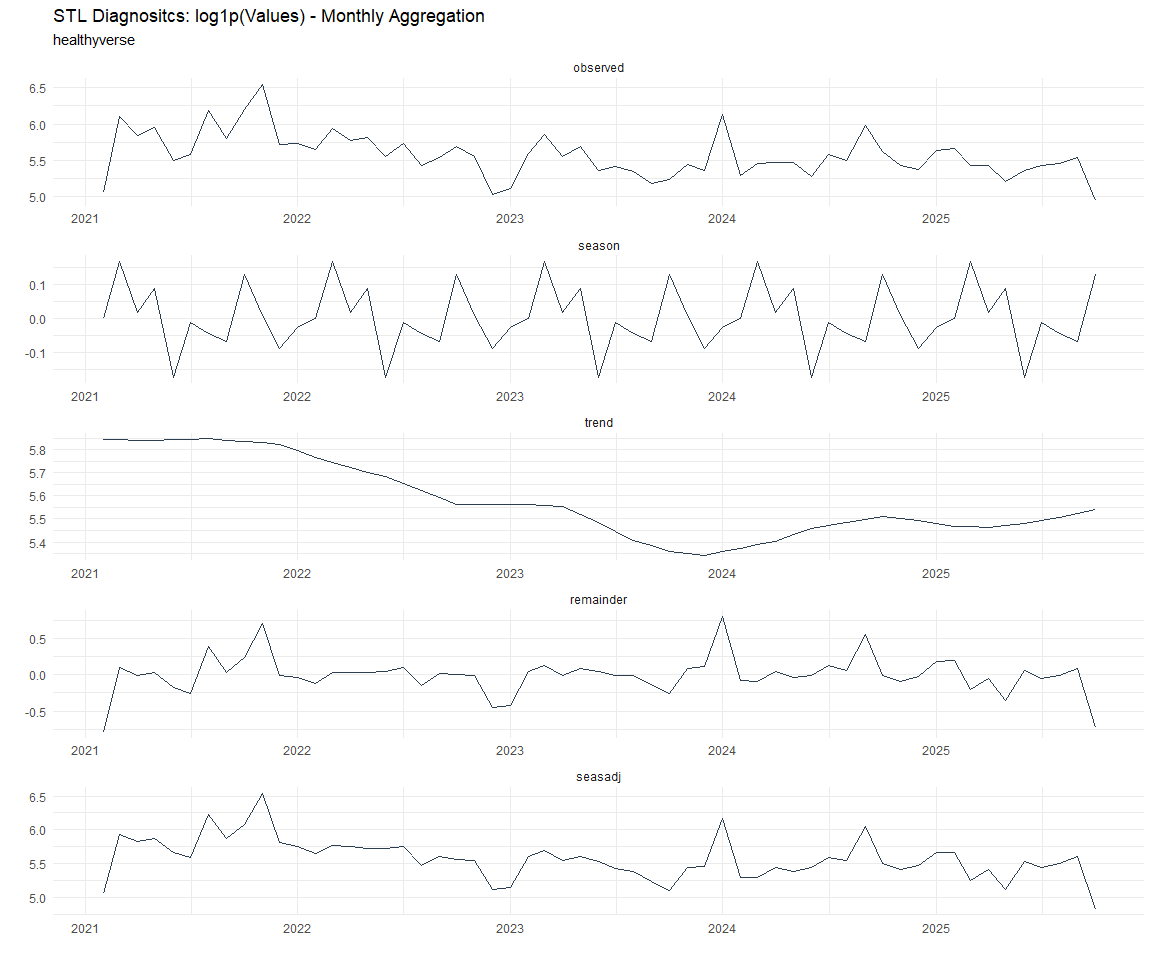
[[8]]
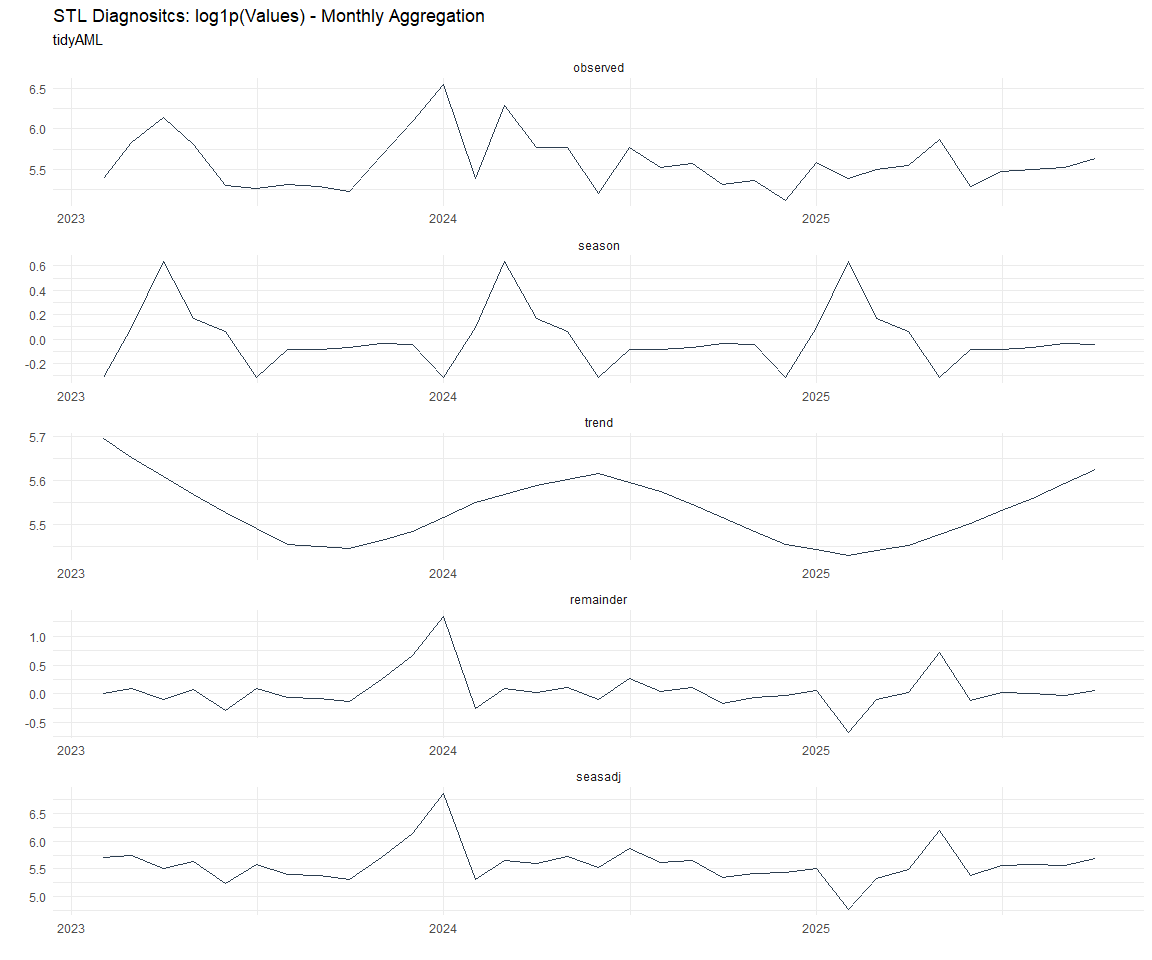
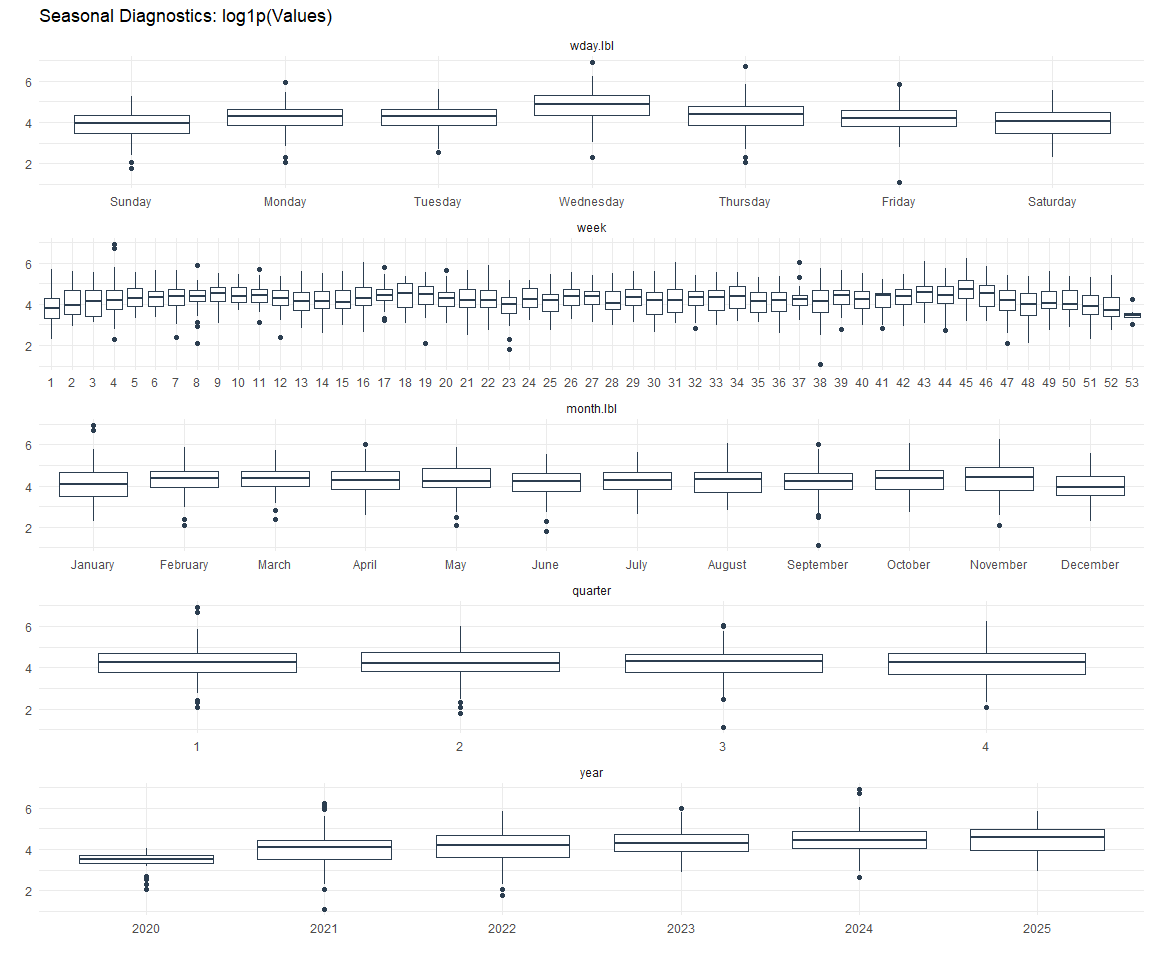
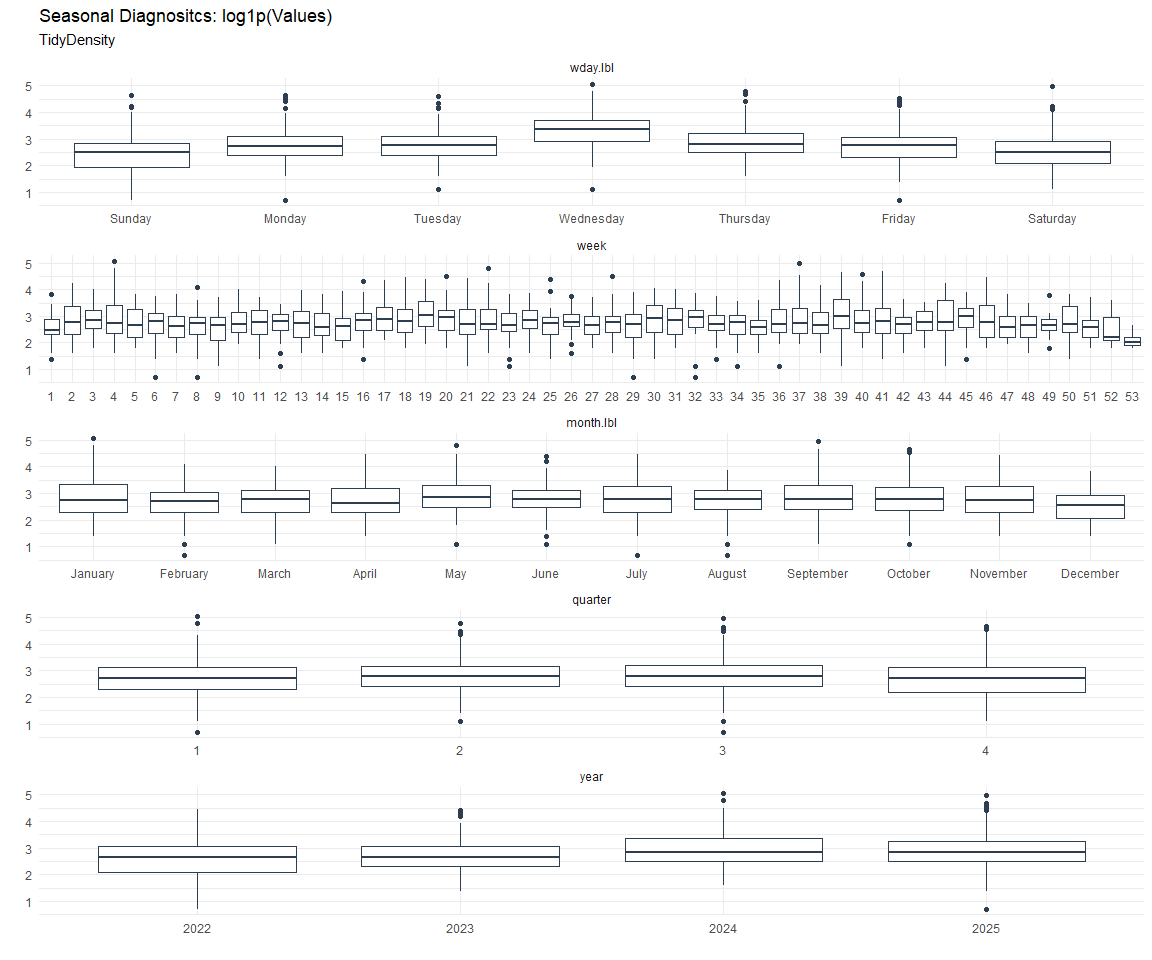
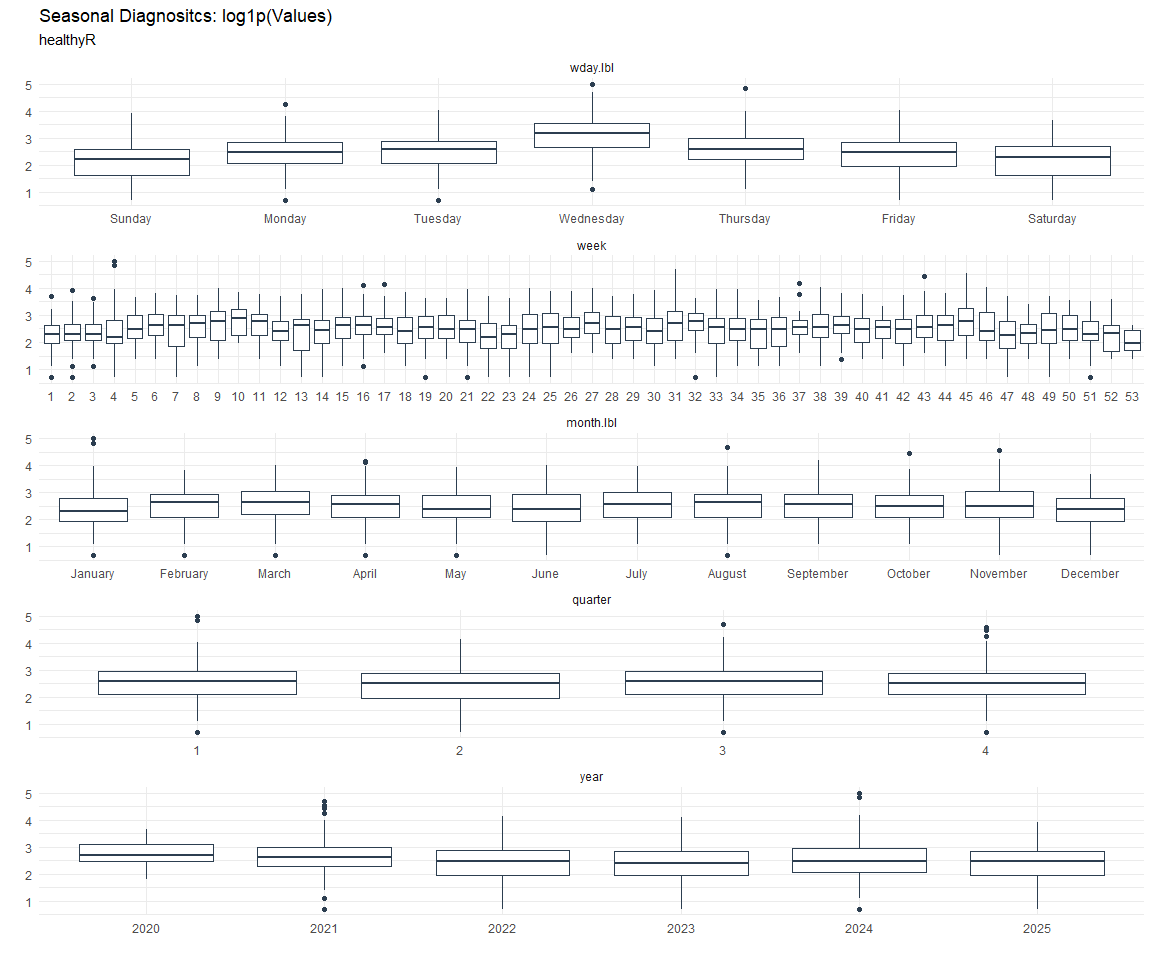
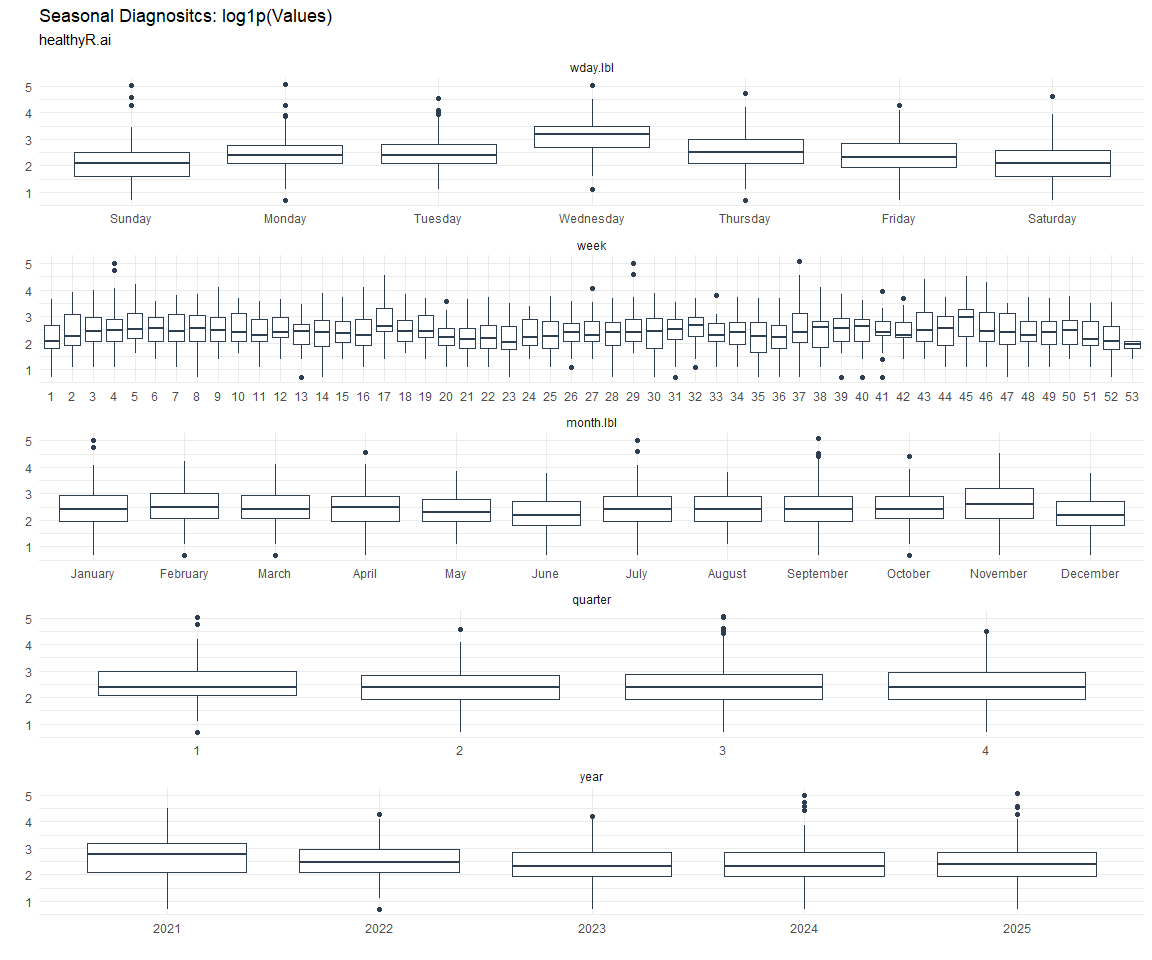
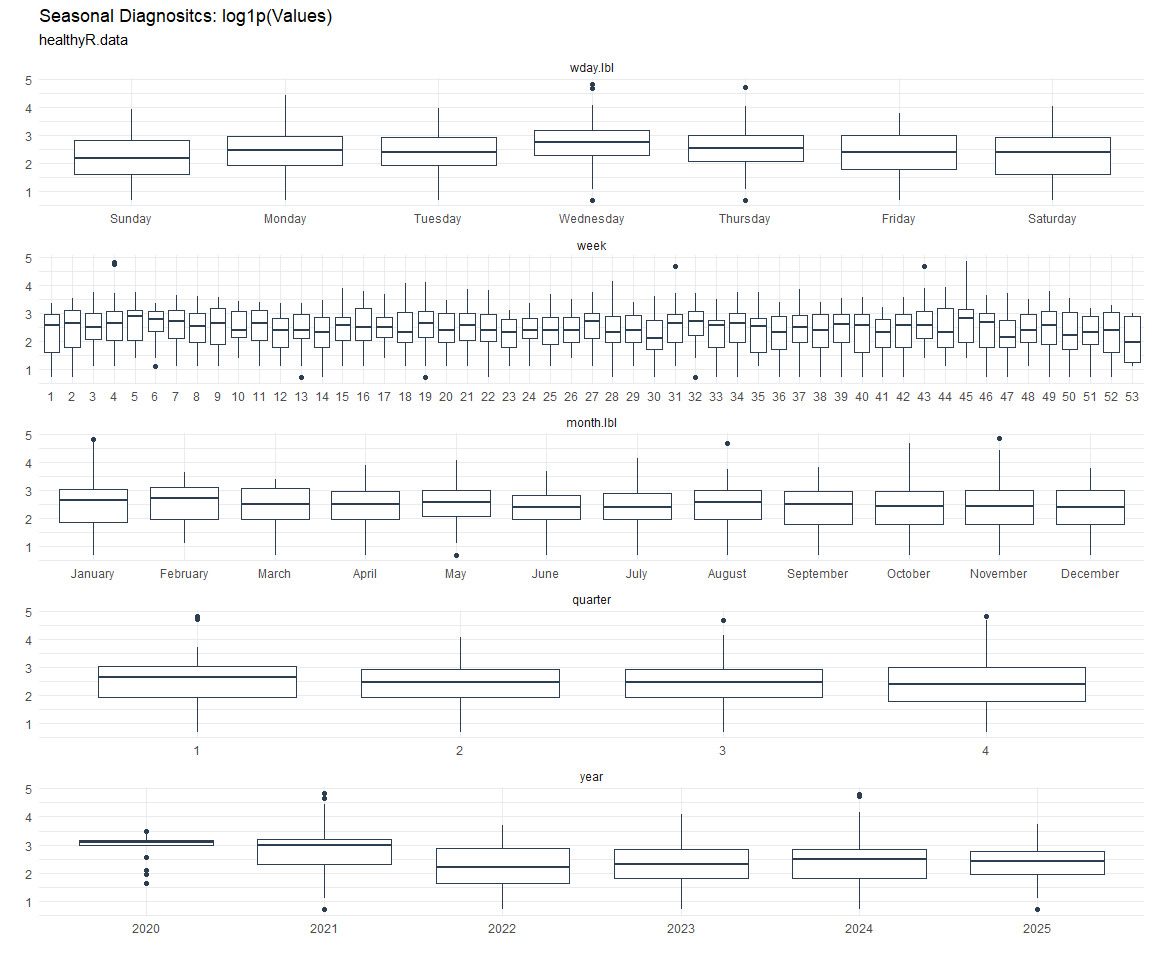
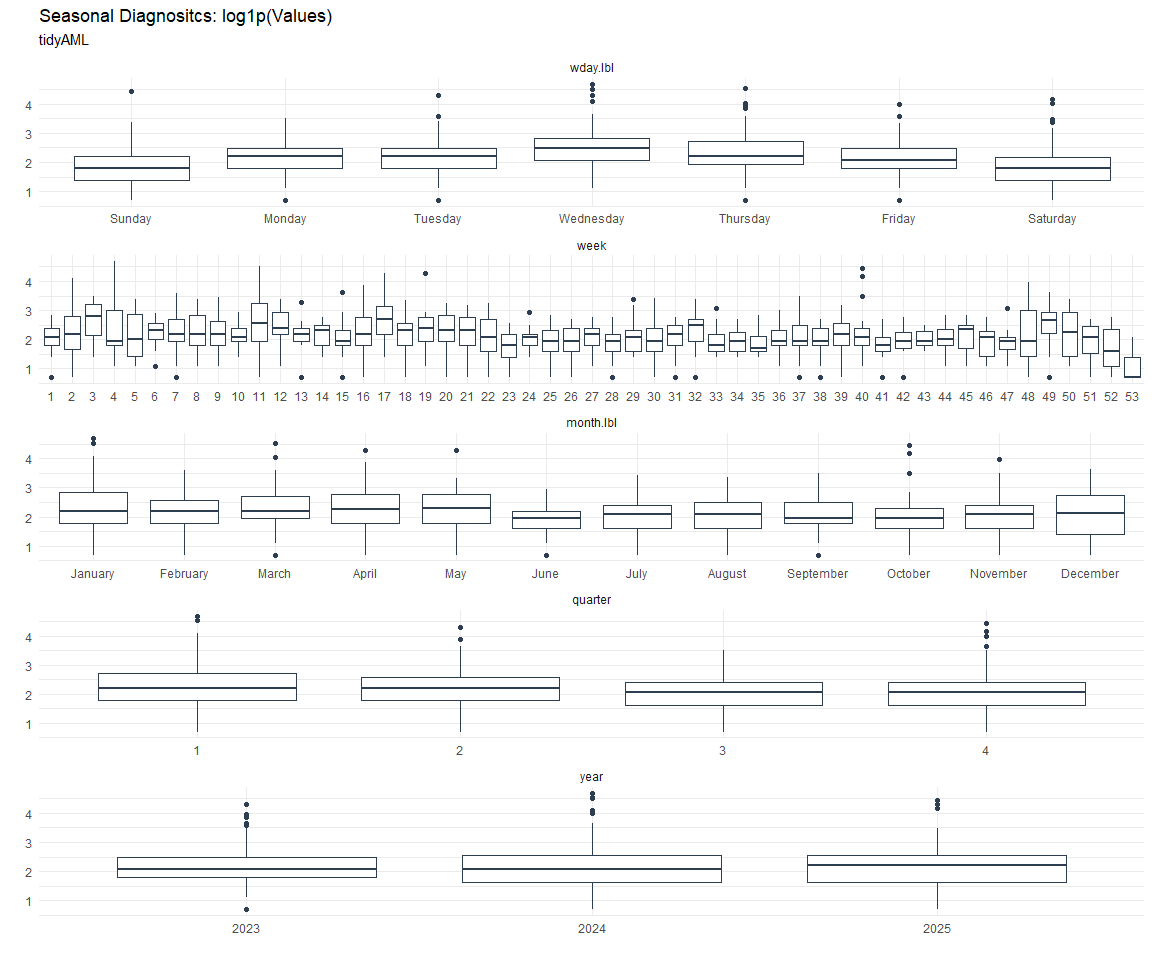
Seasonal Diagnostics:
[[1]]
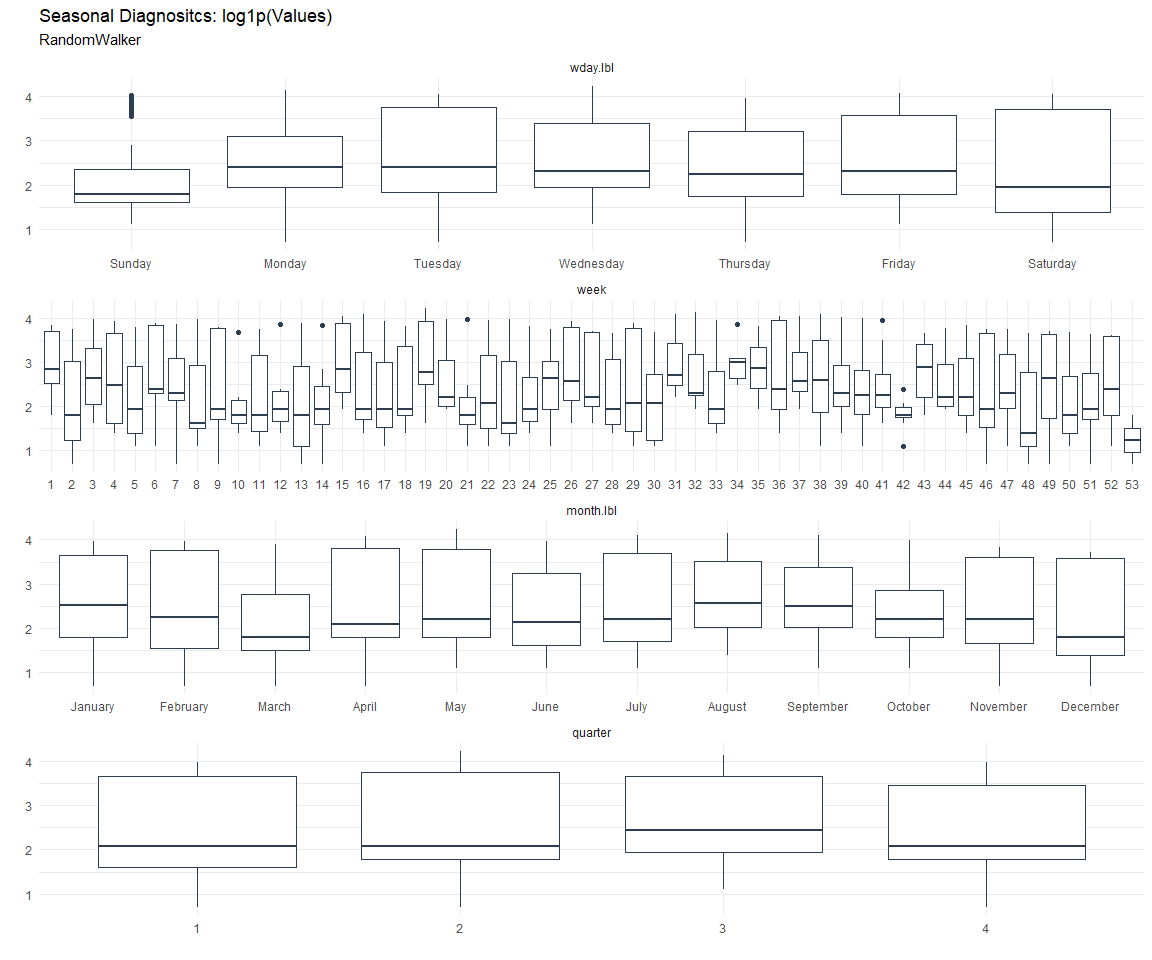
[[2]]
[[3]]
[[4]]
[[5]]
[[6]]
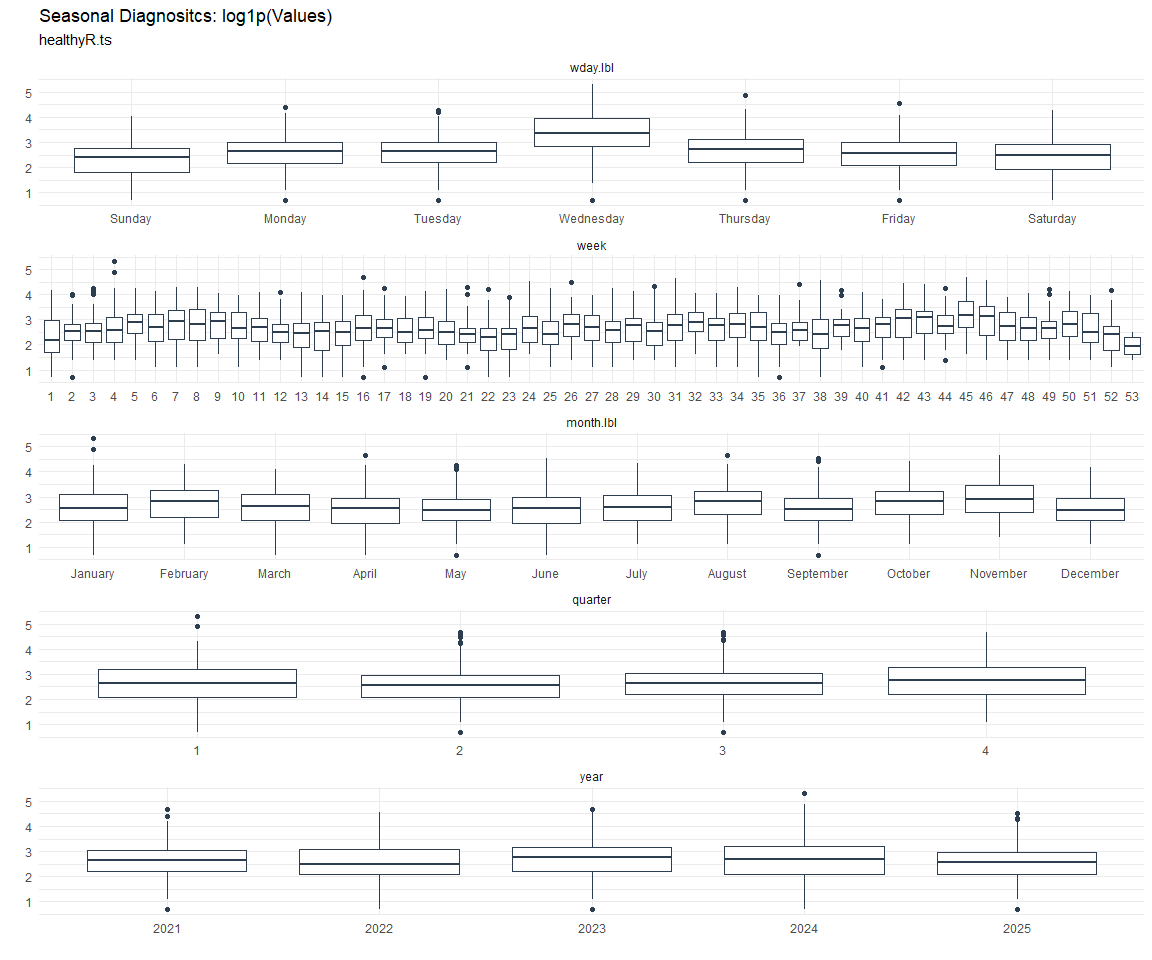
[[7]]
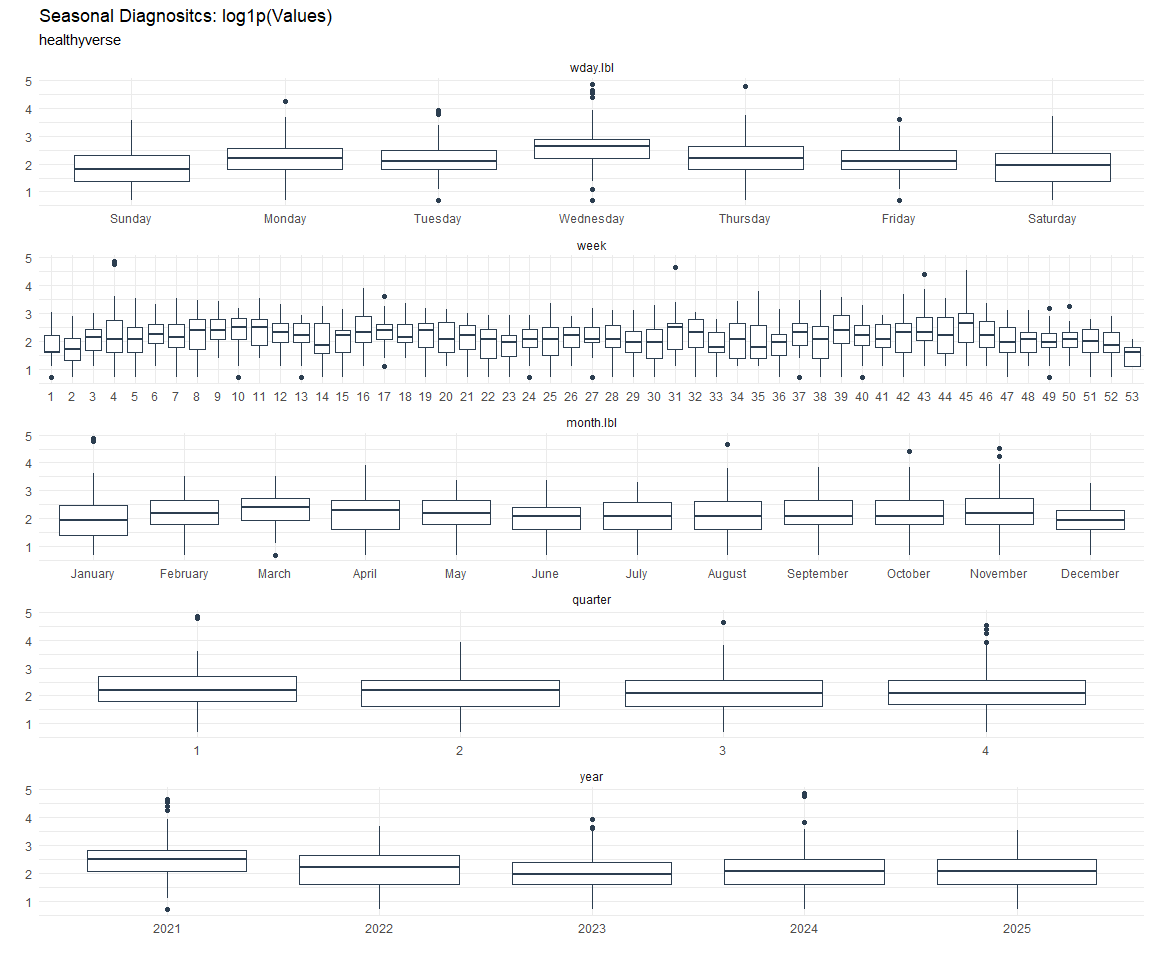
[[8]]
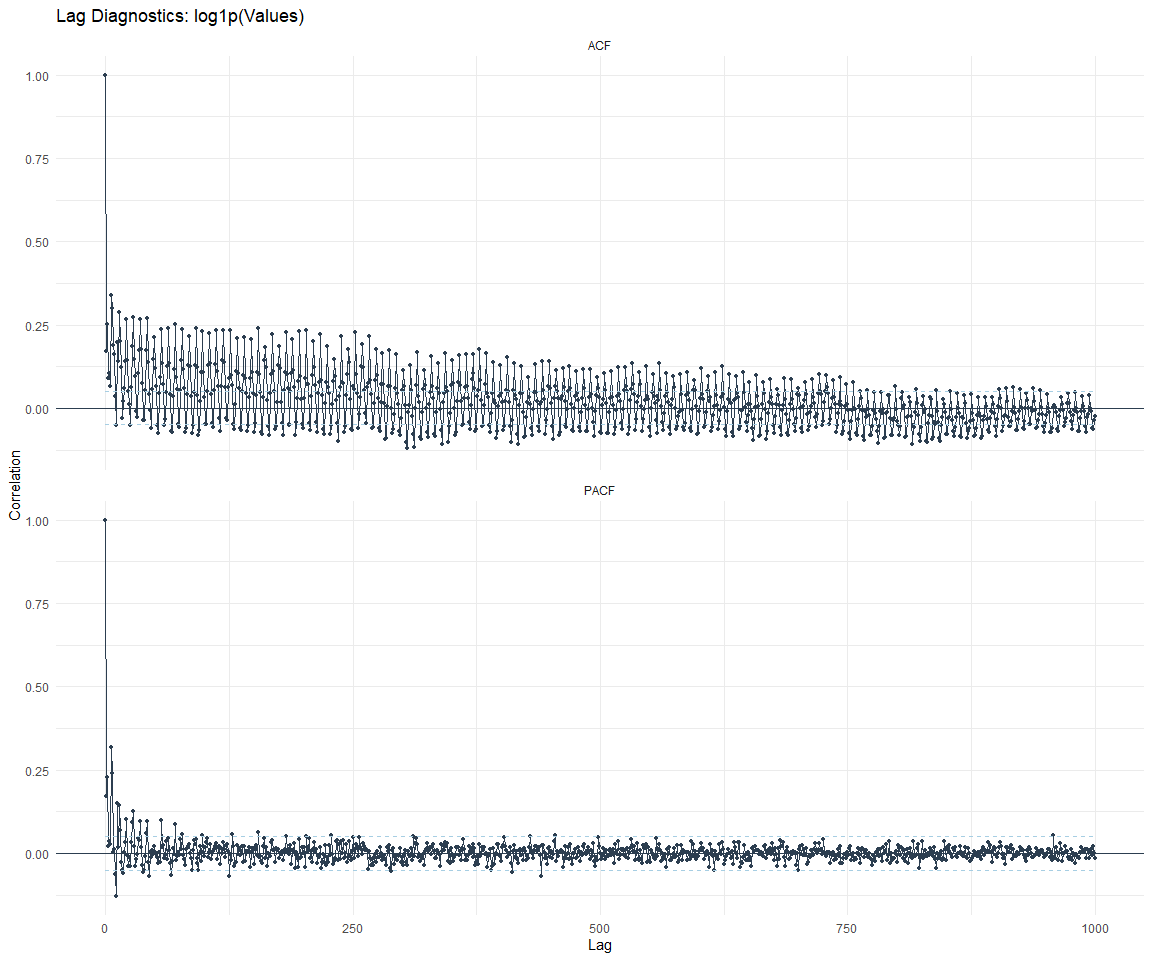
ACF and PACF Diagnostics:
[[1]]
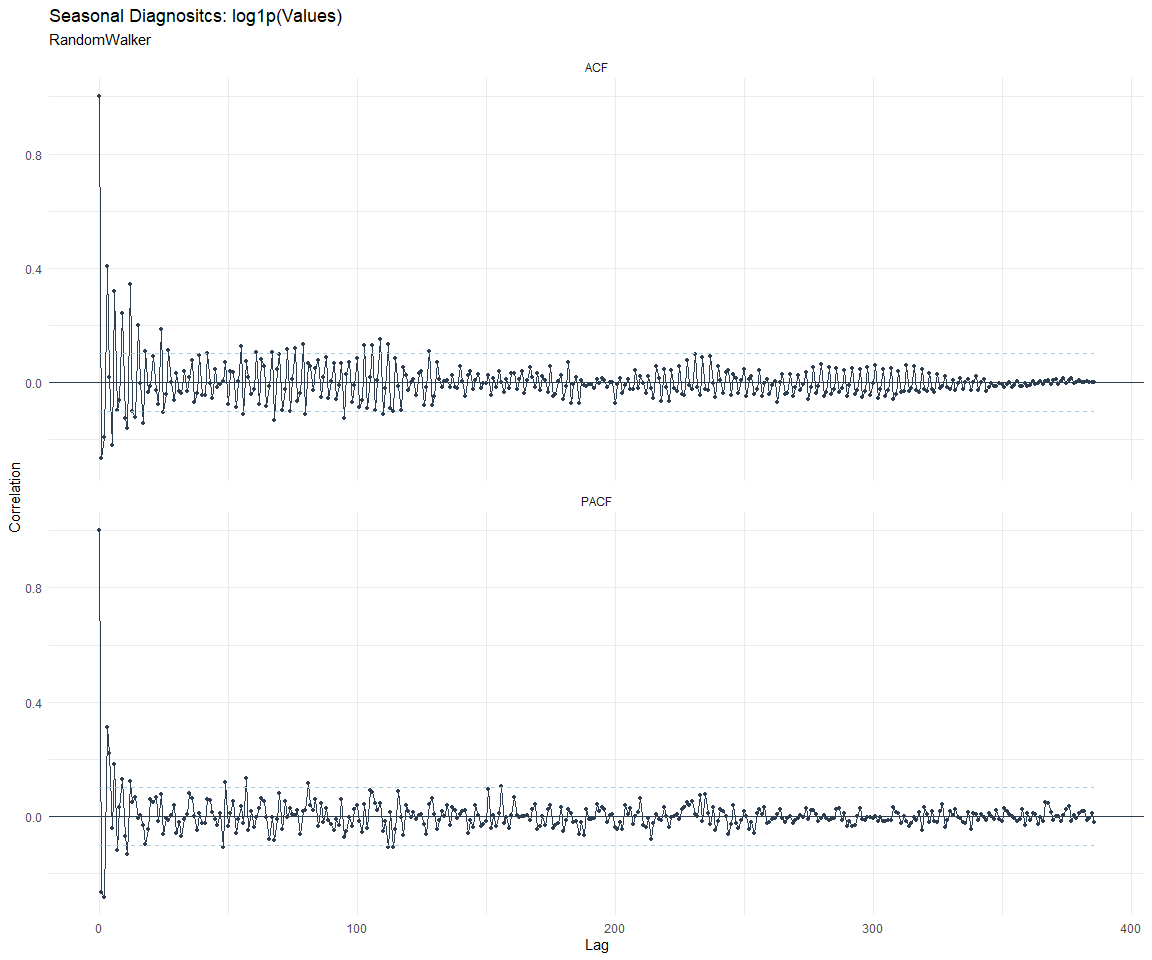
[[2]]
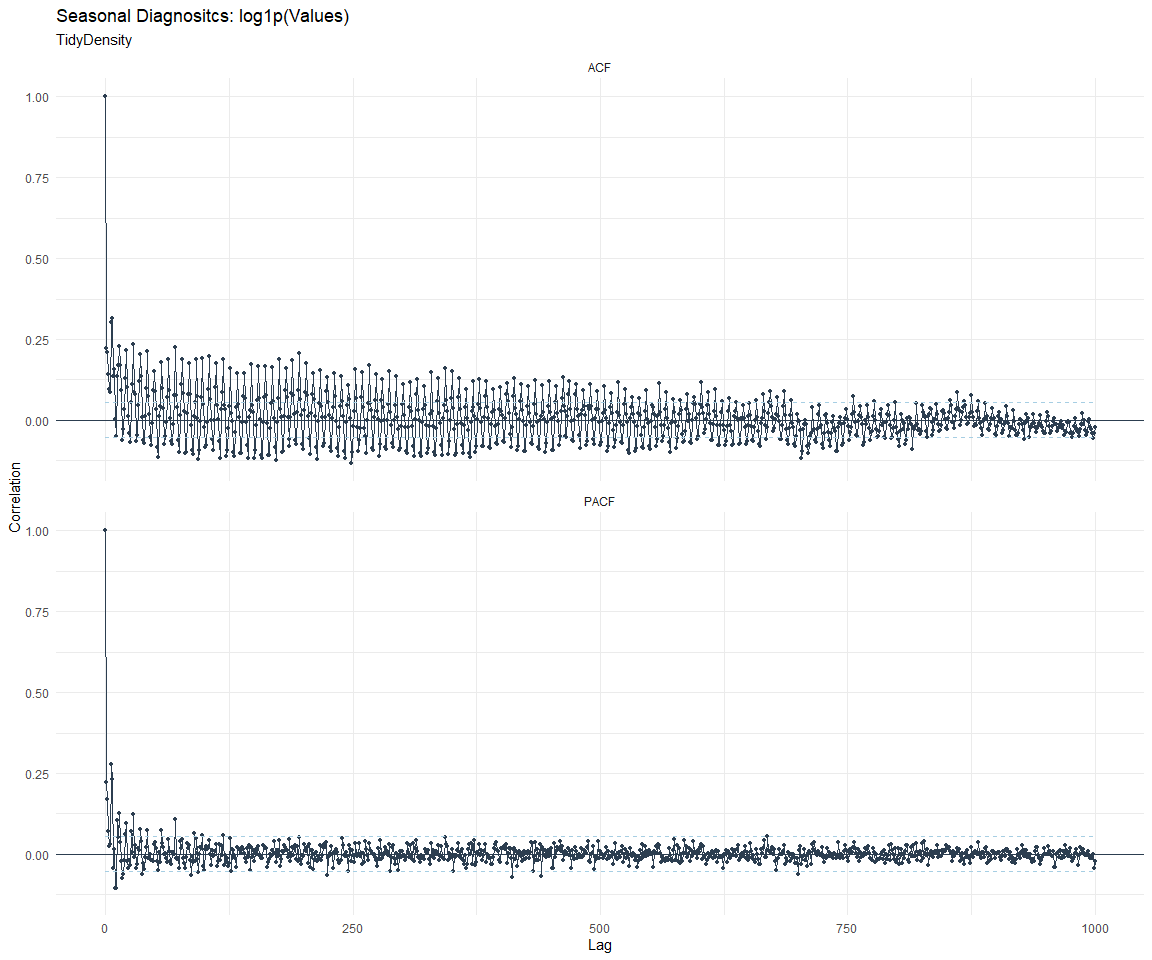
[[3]]
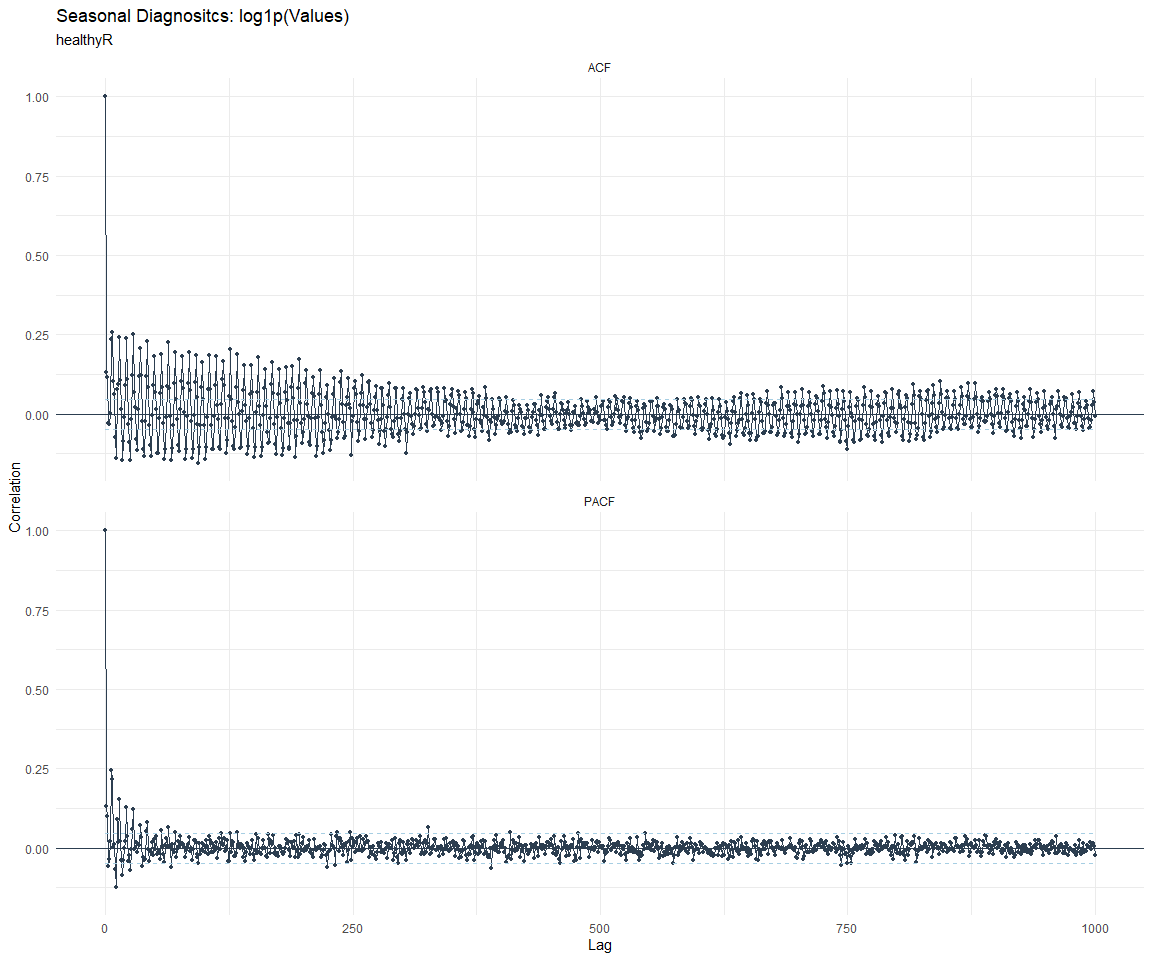
[[4]]
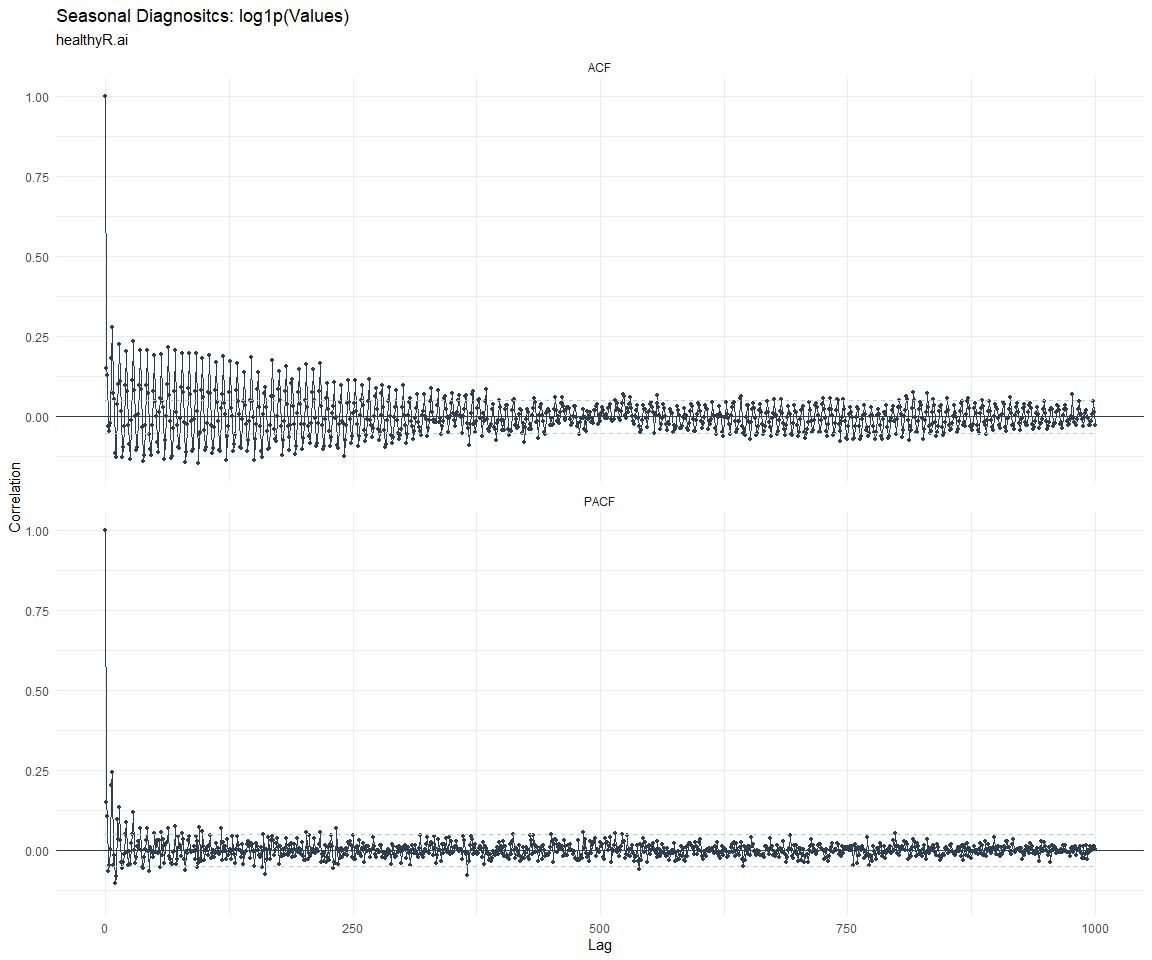
[[5]]
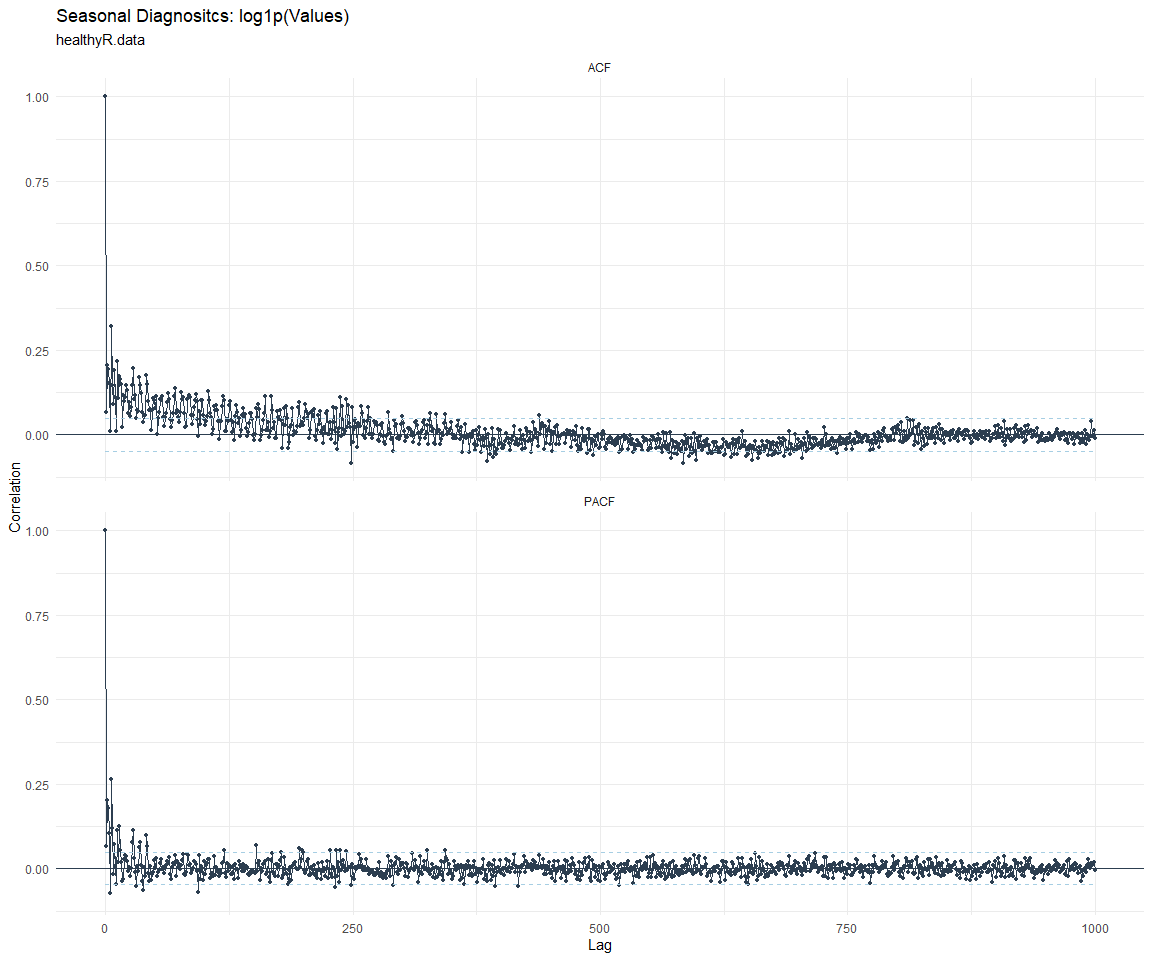
[[6]]
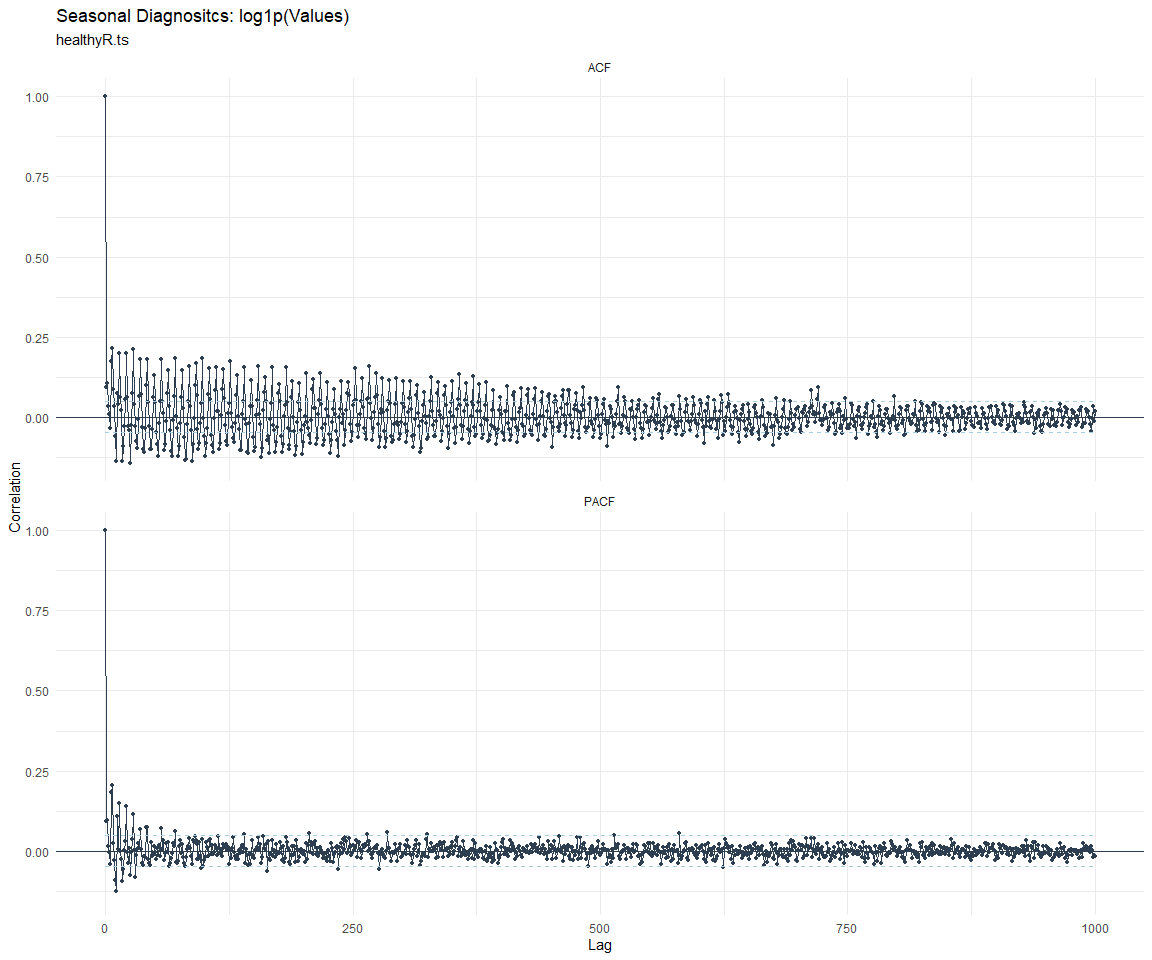
[[7]]
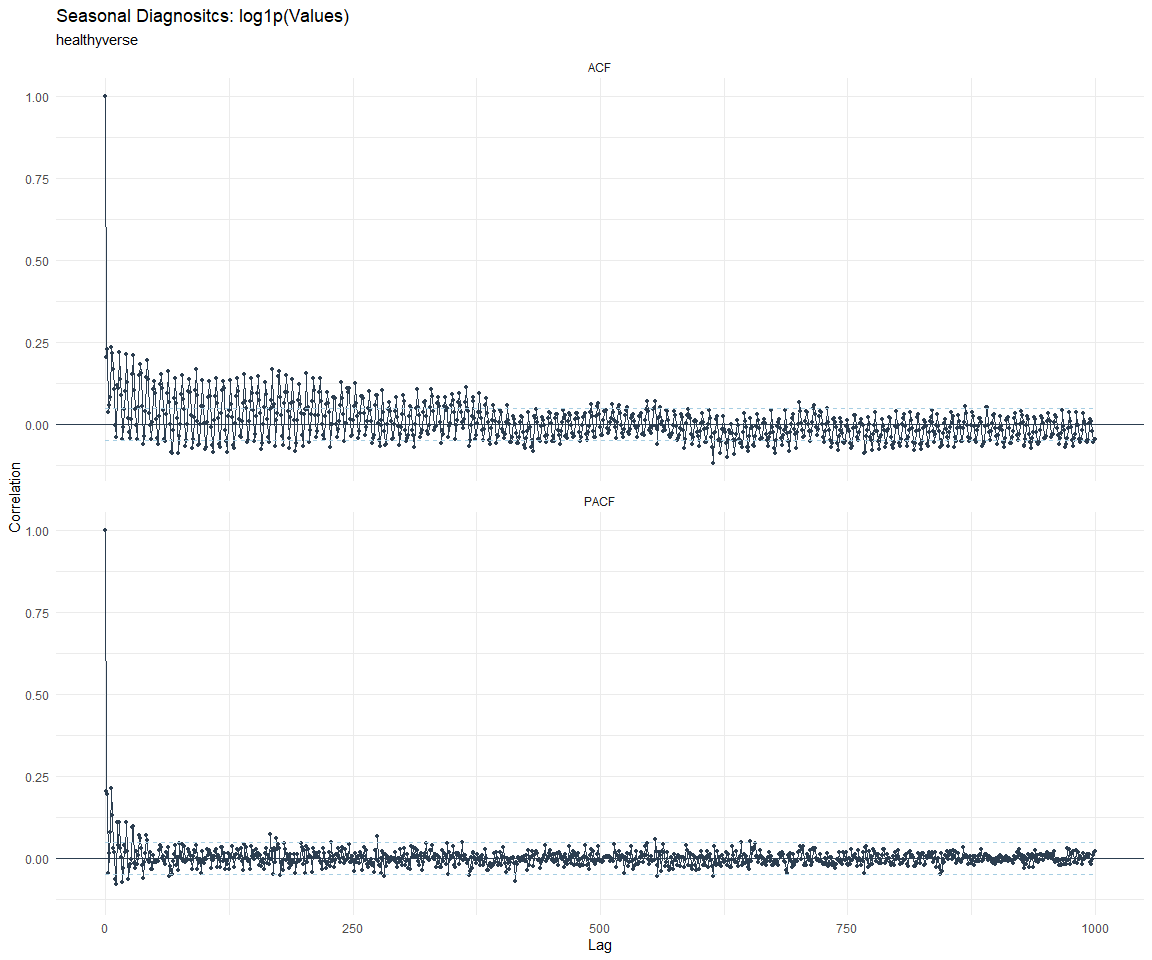
[[8]]
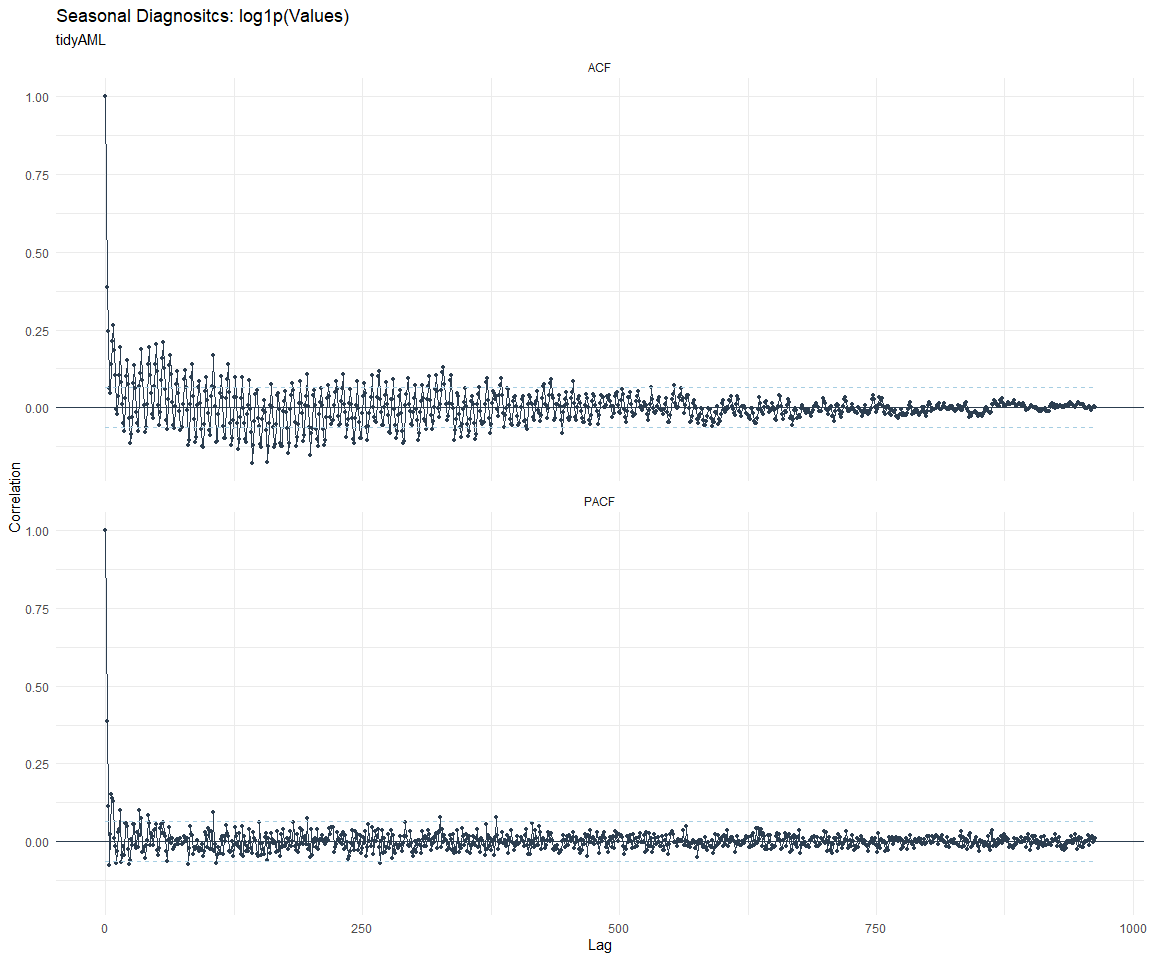
Feature Engineering
Now that we have our basic data and a shot of what it looks like, let’s
add some features to our data which can be very helpful in modeling.
Lets start by making a tibble that is aggregated by the day and
package, as we are going to be interested in forecasting the next 4
weeks or 28 days for each package. First lets get our base data.
Call:
stats::lm(formula = .formula, data = df)
Residuals:
Min 1Q Median 3Q Max
-150.74 -37.72 -11.55 27.91 827.34
Coefficients:
Estimate Std. Error
(Intercept) -1.697e+02 5.338e+01
date 1.063e-02 2.824e-03
lag(value, 1) 8.786e-02 2.251e-02
lag(value, 7) 7.278e-02 2.317e-02
lag(value, 14) 6.583e-02 2.305e-02
lag(value, 21) 8.661e-02 2.313e-02
lag(value, 28) 8.325e-02 2.323e-02
lag(value, 35) 4.485e-02 2.328e-02
lag(value, 42) 6.149e-02 2.340e-02
lag(value, 49) 7.644e-02 2.332e-02
month(date, label = TRUE).L -8.944e+00 4.749e+00
month(date, label = TRUE).Q -7.362e-01 4.773e+00
month(date, label = TRUE).C -1.527e+01 4.753e+00
month(date, label = TRUE)^4 -8.171e+00 4.792e+00
month(date, label = TRUE)^5 -4.938e+00 4.784e+00
month(date, label = TRUE)^6 -9.568e-01 4.791e+00
month(date, label = TRUE)^7 -3.451e+00 4.762e+00
month(date, label = TRUE)^8 -4.638e+00 4.743e+00
month(date, label = TRUE)^9 2.512e+00 4.753e+00
month(date, label = TRUE)^10 1.751e+00 4.831e+00
month(date, label = TRUE)^11 -4.715e+00 4.879e+00
fourier_vec(date, type = "sin", K = 1, period = 7) -1.091e+01 2.146e+00
fourier_vec(date, type = "cos", K = 1, period = 7) 7.575e+00 2.209e+00
t value Pr(>|t|)
(Intercept) -3.179 0.001503 **
date 3.765 0.000172 ***
lag(value, 1) 3.903 9.82e-05 ***
lag(value, 7) 3.141 0.001707 **
lag(value, 14) 2.856 0.004334 **
lag(value, 21) 3.744 0.000187 ***
lag(value, 28) 3.584 0.000347 ***
lag(value, 35) 1.927 0.054149 .
lag(value, 42) 2.628 0.008662 **
lag(value, 49) 3.278 0.001065 **
month(date, label = TRUE).L -1.884 0.059782 .
month(date, label = TRUE).Q -0.154 0.877434
month(date, label = TRUE).C -3.213 0.001337 **
month(date, label = TRUE)^4 -1.705 0.088301 .
month(date, label = TRUE)^5 -1.032 0.302022
month(date, label = TRUE)^6 -0.200 0.841741
month(date, label = TRUE)^7 -0.725 0.468689
month(date, label = TRUE)^8 -0.978 0.328234
month(date, label = TRUE)^9 0.529 0.597154
month(date, label = TRUE)^10 0.362 0.717094
month(date, label = TRUE)^11 -0.966 0.333949
fourier_vec(date, type = "sin", K = 1, period = 7) -5.085 4.03e-07 ***
fourier_vec(date, type = "cos", K = 1, period = 7) 3.429 0.000618 ***
---
Signif. codes: 0 '***' 0.001 '**' 0.01 '*' 0.05 '.' 0.1 ' ' 1
Residual standard error: 60.08 on 1903 degrees of freedom
(49 observations deleted due to missingness)
Multiple R-squared: 0.2098, Adjusted R-squared: 0.2006
F-statistic: 22.96 on 22 and 1903 DF, p-value: < 2.2e-16
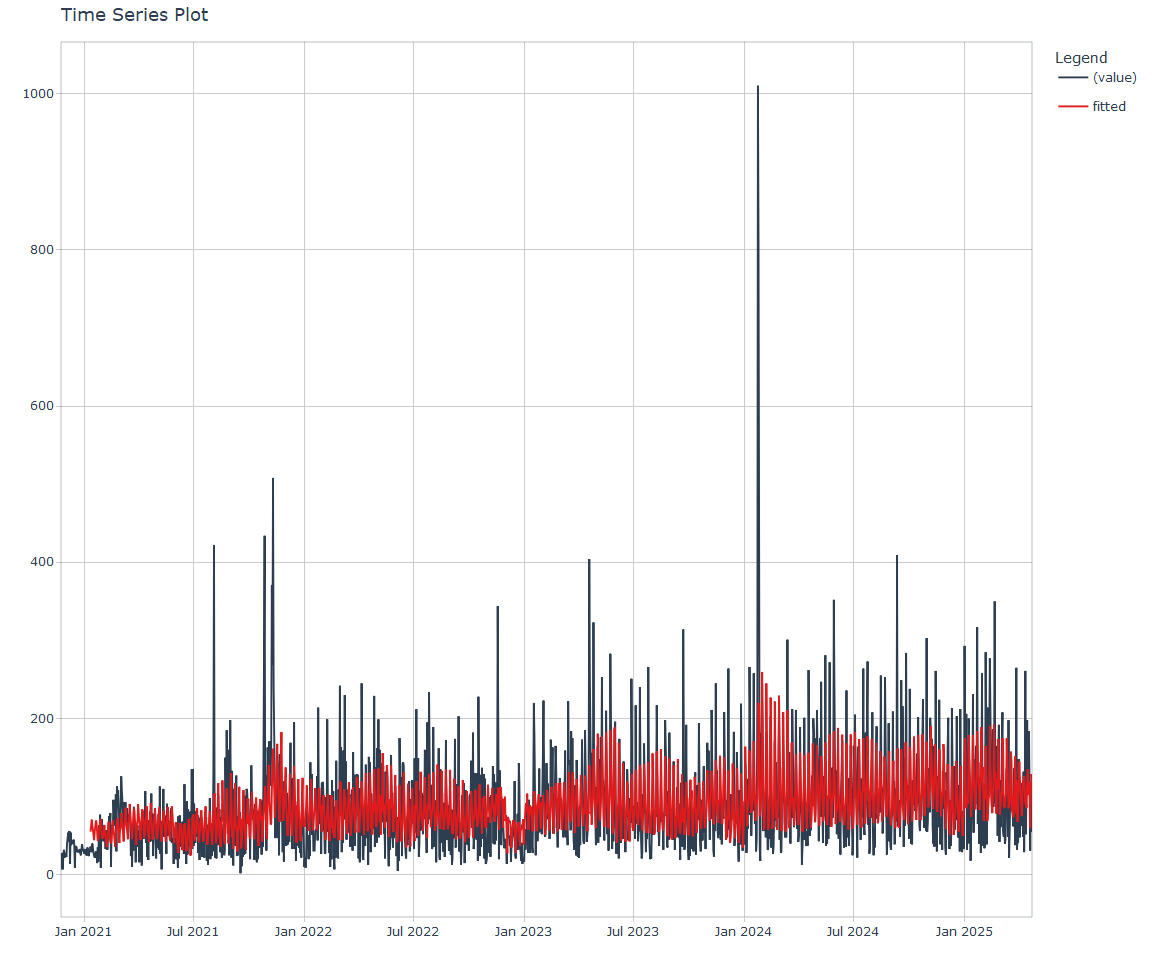
NNS Forecasting
This is something I have been wanting to try for a while. The NNS
package is a great package for forecasting time series data.
library(NNS)
data_list <- base_data |>
select(package, value) |>
group_split(package)
data_list |>
imap(
\(x, idx) {
obj <- x
x <- obj |> pull(value) |> tail(7*52)
train_set_size <- length(x) - 56
pkg <- obj |> pluck(1) |> unique()
# sf <- NNS.seas(x, modulo = 7, plot = FALSE)$periods
seas <- t(
sapply(
1:25,
function(i) c(
i,
sqrt(
mean((
NNS.ARMA(x,
h = 28,
training.set = train_set_size,
method = "lin",
seasonal.factor = i,
plot=FALSE
) - tail(x, 28)) ^ 2)))
)
)
colnames(seas) <- c("Period", "RMSE")
sf <- seas[which.min(seas[, 2]), 1]
cat(paste0("Package: ", pkg, "\n"))
NNS.ARMA.optim(
variable = x,
h = 28,
training.set = train_set_size,
#seasonal.factor = seq(12, 60, 7),
seasonal.factor = sf,
pred.int = 0.95,
plot = TRUE
)
title(
sub = paste0("\n",
"Package: ", pkg, " - NNS Optimization")
)
}
)
Package: healthyR
[1] "CURRNET METHOD: lin"
[1] "COPY LATEST PARAMETERS DIRECTLY FOR NNS.ARMA() IF ERROR:"
[1] "NNS.ARMA(... method = 'lin' , seasonal.factor = c( 18 ) ...)"
[1] "CURRENT lin OBJECTIVE FUNCTION = 21.6150670259139"
[1] "BEST method = 'lin' PATH MEMBER = c( 18 )"
[1] "BEST lin OBJECTIVE FUNCTION = 21.6150670259139"
[1] "CURRNET METHOD: nonlin"
[1] "COPY LATEST PARAMETERS DIRECTLY FOR NNS.ARMA() IF ERROR:"
[1] "NNS.ARMA(... method = 'nonlin' , seasonal.factor = c( 18 ) ...)"
[1] "CURRENT nonlin OBJECTIVE FUNCTION = 10.2932340733006"
[1] "BEST method = 'nonlin' PATH MEMBER = c( 18 )"
[1] "BEST nonlin OBJECTIVE FUNCTION = 10.2932340733006"
[1] "CURRNET METHOD: both"
[1] "COPY LATEST PARAMETERS DIRECTLY FOR NNS.ARMA() IF ERROR:"
[1] "NNS.ARMA(... method = 'both' , seasonal.factor = c( 18 ) ...)"
[1] "CURRENT both OBJECTIVE FUNCTION = 11.8605224292463"
[1] "BEST method = 'both' PATH MEMBER = c( 18 )"
[1] "BEST both OBJECTIVE FUNCTION = 11.8605224292463"
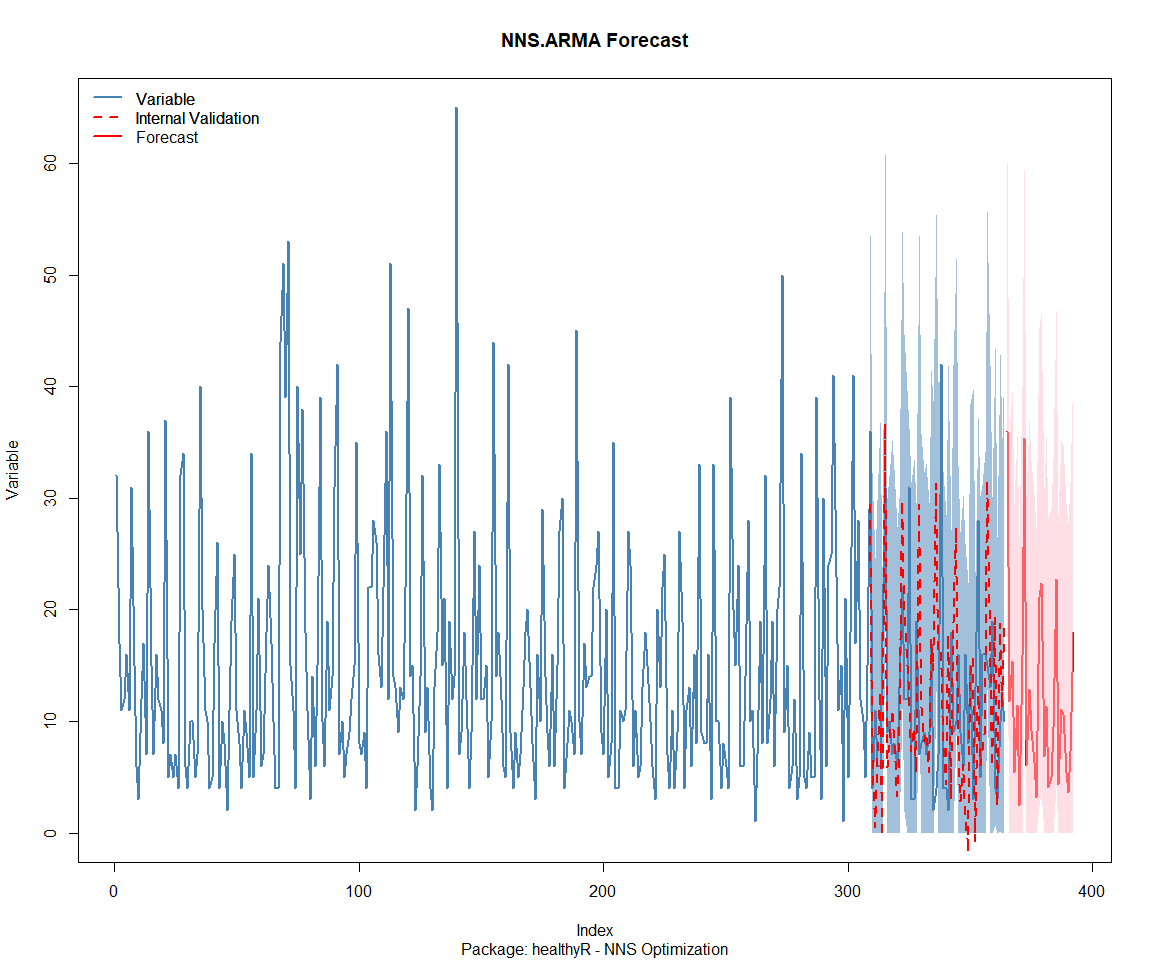
Package: healthyR.ai
[1] "CURRNET METHOD: lin"
[1] "COPY LATEST PARAMETERS DIRECTLY FOR NNS.ARMA() IF ERROR:"
[1] "NNS.ARMA(... method = 'lin' , seasonal.factor = c( 24 ) ...)"
[1] "CURRENT lin OBJECTIVE FUNCTION = 9.44069345491946"
[1] "BEST method = 'lin' PATH MEMBER = c( 24 )"
[1] "BEST lin OBJECTIVE FUNCTION = 9.44069345491946"
[1] "CURRNET METHOD: nonlin"
[1] "COPY LATEST PARAMETERS DIRECTLY FOR NNS.ARMA() IF ERROR:"
[1] "NNS.ARMA(... method = 'nonlin' , seasonal.factor = c( 24 ) ...)"
[1] "CURRENT nonlin OBJECTIVE FUNCTION = 15.0513569793399"
[1] "BEST method = 'nonlin' PATH MEMBER = c( 24 )"
[1] "BEST nonlin OBJECTIVE FUNCTION = 15.0513569793399"
[1] "CURRNET METHOD: both"
[1] "COPY LATEST PARAMETERS DIRECTLY FOR NNS.ARMA() IF ERROR:"
[1] "NNS.ARMA(... method = 'both' , seasonal.factor = c( 24 ) ...)"
[1] "CURRENT both OBJECTIVE FUNCTION = 12.3828547130066"
[1] "BEST method = 'both' PATH MEMBER = c( 24 )"
[1] "BEST both OBJECTIVE FUNCTION = 12.3828547130066"
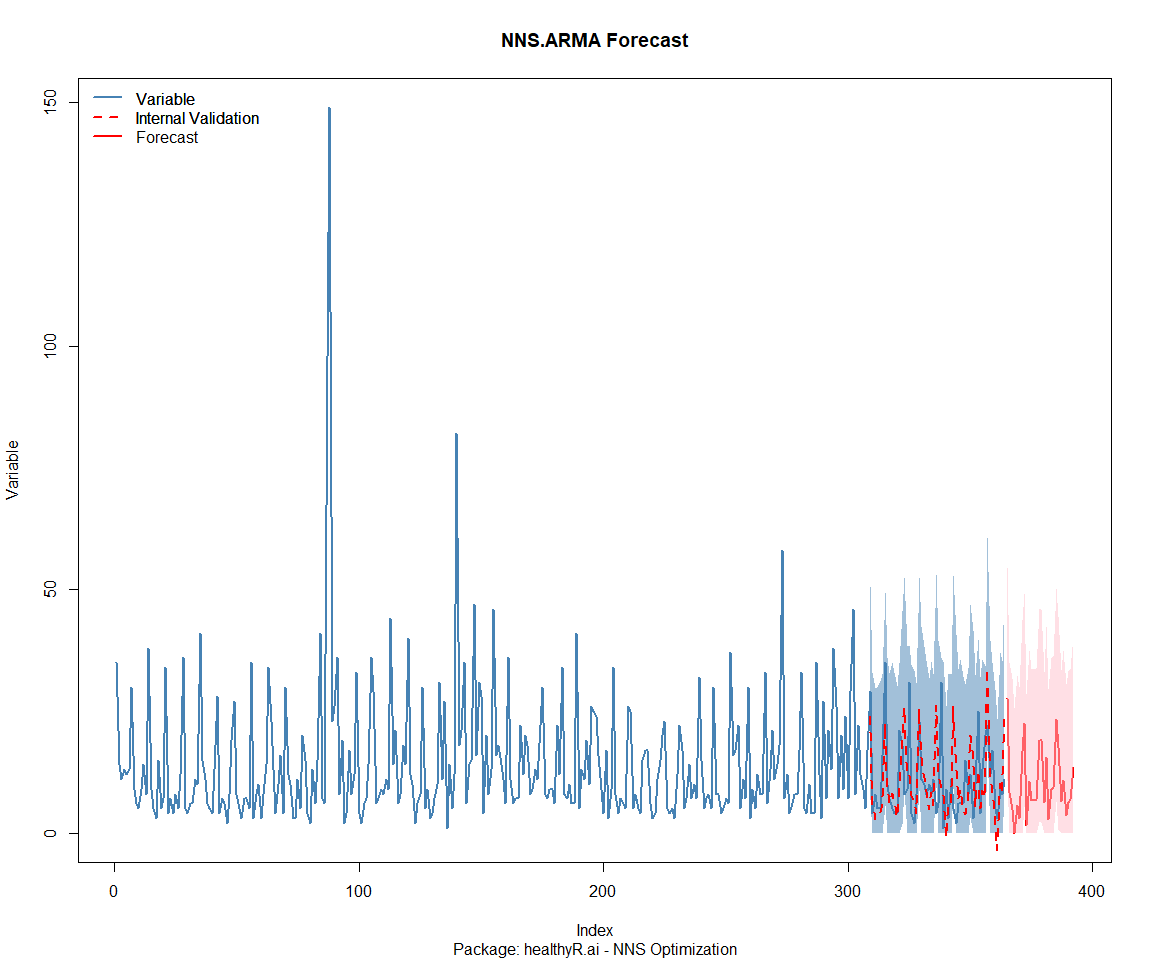
Package: healthyR.data
[1] "CURRNET METHOD: lin"
[1] "COPY LATEST PARAMETERS DIRECTLY FOR NNS.ARMA() IF ERROR:"
[1] "NNS.ARMA(... method = 'lin' , seasonal.factor = c( 19 ) ...)"
[1] "CURRENT lin OBJECTIVE FUNCTION = 25.7022527384029"
[1] "BEST method = 'lin' PATH MEMBER = c( 19 )"
[1] "BEST lin OBJECTIVE FUNCTION = 25.7022527384029"
[1] "CURRNET METHOD: nonlin"
[1] "COPY LATEST PARAMETERS DIRECTLY FOR NNS.ARMA() IF ERROR:"
[1] "NNS.ARMA(... method = 'nonlin' , seasonal.factor = c( 19 ) ...)"
[1] "CURRENT nonlin OBJECTIVE FUNCTION = 6.25705611400315"
[1] "BEST method = 'nonlin' PATH MEMBER = c( 19 )"
[1] "BEST nonlin OBJECTIVE FUNCTION = 6.25705611400315"
[1] "CURRNET METHOD: both"
[1] "COPY LATEST PARAMETERS DIRECTLY FOR NNS.ARMA() IF ERROR:"
[1] "NNS.ARMA(... method = 'both' , seasonal.factor = c( 19 ) ...)"
[1] "CURRENT both OBJECTIVE FUNCTION = 10.3498705763747"
[1] "BEST method = 'both' PATH MEMBER = c( 19 )"
[1] "BEST both OBJECTIVE FUNCTION = 10.3498705763747"
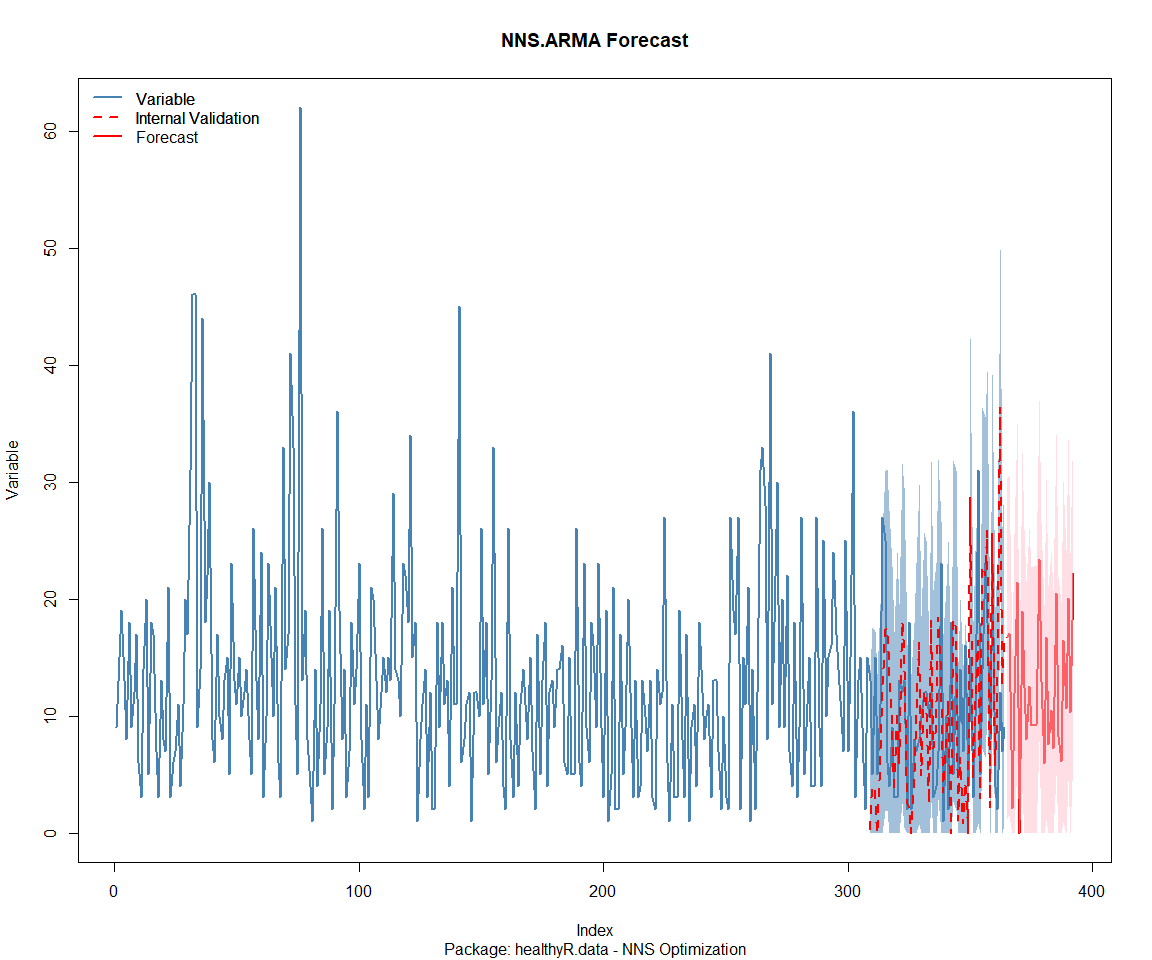
Package: healthyR.ts
[1] "CURRNET METHOD: lin"
[1] "COPY LATEST PARAMETERS DIRECTLY FOR NNS.ARMA() IF ERROR:"
[1] "NNS.ARMA(... method = 'lin' , seasonal.factor = c( 17 ) ...)"
[1] "CURRENT lin OBJECTIVE FUNCTION = 11.1209157764124"
[1] "BEST method = 'lin' PATH MEMBER = c( 17 )"
[1] "BEST lin OBJECTIVE FUNCTION = 11.1209157764124"
[1] "CURRNET METHOD: nonlin"
[1] "COPY LATEST PARAMETERS DIRECTLY FOR NNS.ARMA() IF ERROR:"
[1] "NNS.ARMA(... method = 'nonlin' , seasonal.factor = c( 17 ) ...)"
[1] "CURRENT nonlin OBJECTIVE FUNCTION = 11.4602249137556"
[1] "BEST method = 'nonlin' PATH MEMBER = c( 17 )"
[1] "BEST nonlin OBJECTIVE FUNCTION = 11.4602249137556"
[1] "CURRNET METHOD: both"
[1] "COPY LATEST PARAMETERS DIRECTLY FOR NNS.ARMA() IF ERROR:"
[1] "NNS.ARMA(... method = 'both' , seasonal.factor = c( 17 ) ...)"
[1] "CURRENT both OBJECTIVE FUNCTION = 8.98014581385281"
[1] "BEST method = 'both' PATH MEMBER = c( 17 )"
[1] "BEST both OBJECTIVE FUNCTION = 8.98014581385281"
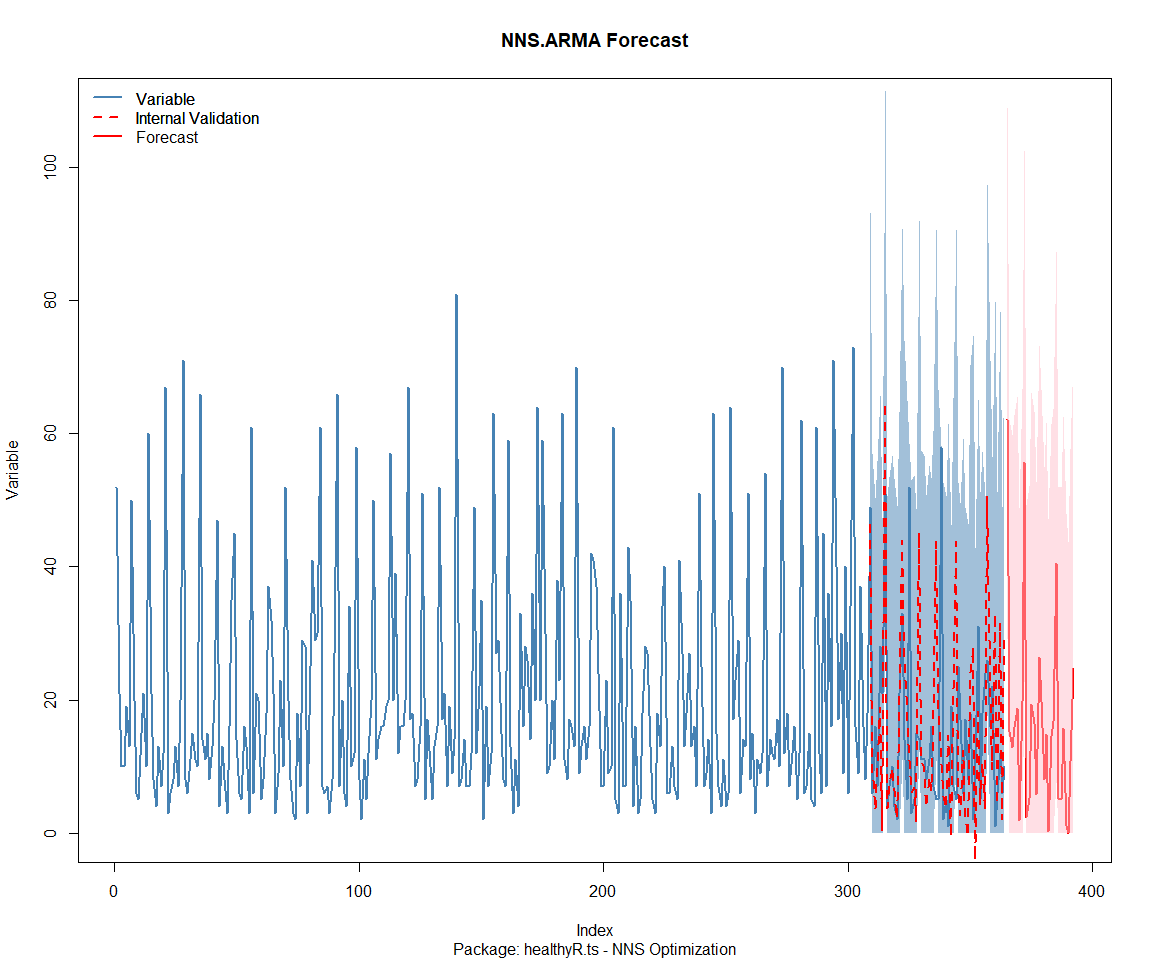
Package: healthyverse
[1] "CURRNET METHOD: lin"
[1] "COPY LATEST PARAMETERS DIRECTLY FOR NNS.ARMA() IF ERROR:"
[1] "NNS.ARMA(... method = 'lin' , seasonal.factor = c( 9 ) ...)"
[1] "CURRENT lin OBJECTIVE FUNCTION = 41.1878734472022"
[1] "BEST method = 'lin' PATH MEMBER = c( 9 )"
[1] "BEST lin OBJECTIVE FUNCTION = 41.1878734472022"
[1] "CURRNET METHOD: nonlin"
[1] "COPY LATEST PARAMETERS DIRECTLY FOR NNS.ARMA() IF ERROR:"
[1] "NNS.ARMA(... method = 'nonlin' , seasonal.factor = c( 9 ) ...)"
[1] "CURRENT nonlin OBJECTIVE FUNCTION = 19.9246283446847"
[1] "BEST method = 'nonlin' PATH MEMBER = c( 9 )"
[1] "BEST nonlin OBJECTIVE FUNCTION = 19.9246283446847"
[1] "CURRNET METHOD: both"
[1] "COPY LATEST PARAMETERS DIRECTLY FOR NNS.ARMA() IF ERROR:"
[1] "NNS.ARMA(... method = 'both' , seasonal.factor = c( 9 ) ...)"
[1] "CURRENT both OBJECTIVE FUNCTION = 20.3439867246216"
[1] "BEST method = 'both' PATH MEMBER = c( 9 )"
[1] "BEST both OBJECTIVE FUNCTION = 20.3439867246216"
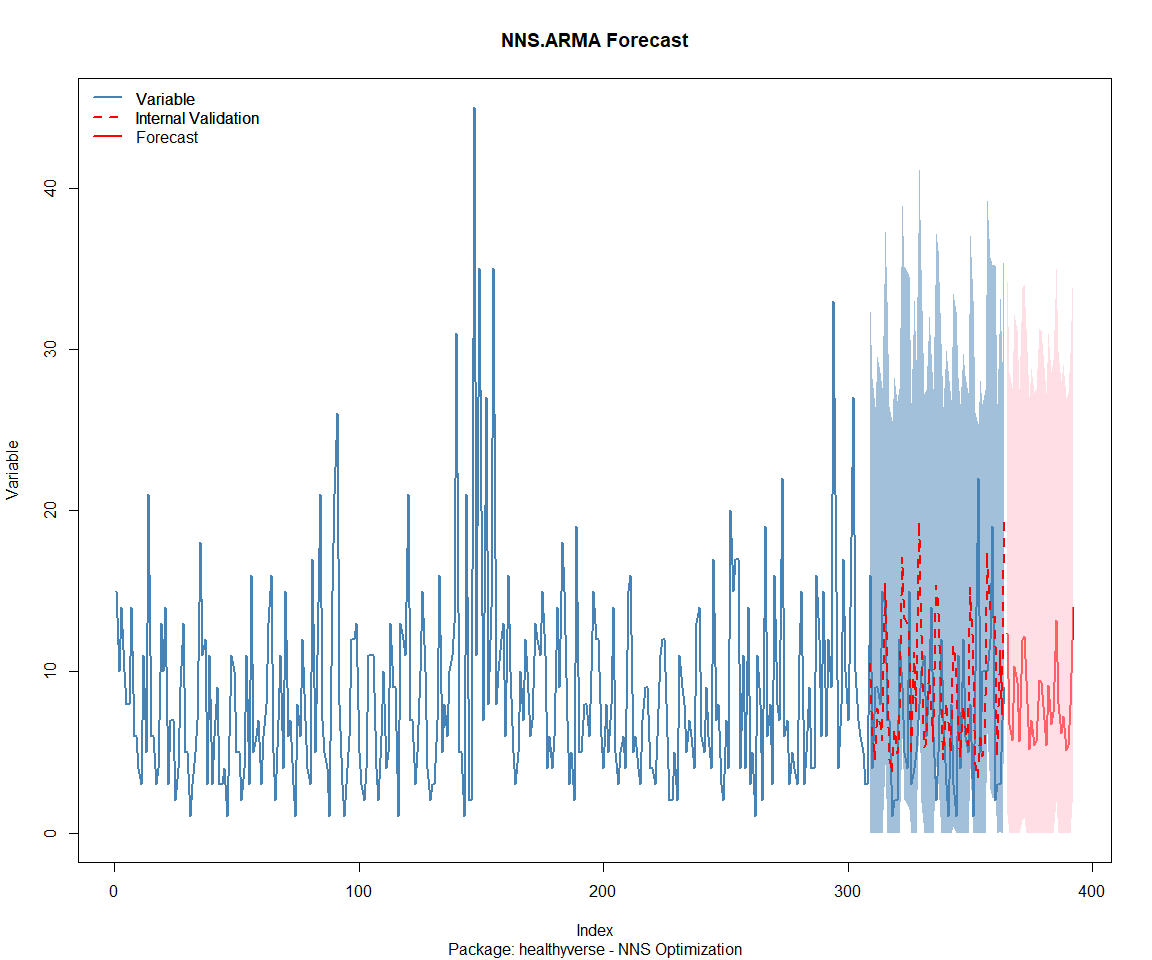
Package: RandomWalker
[1] "CURRNET METHOD: lin"
[1] "COPY LATEST PARAMETERS DIRECTLY FOR NNS.ARMA() IF ERROR:"
[1] "NNS.ARMA(... method = 'lin' , seasonal.factor = c( 17 ) ...)"
[1] "CURRENT lin OBJECTIVE FUNCTION = 4.30329048442896"
[1] "BEST method = 'lin' PATH MEMBER = c( 17 )"
[1] "BEST lin OBJECTIVE FUNCTION = 4.30329048442896"
[1] "CURRNET METHOD: nonlin"
[1] "COPY LATEST PARAMETERS DIRECTLY FOR NNS.ARMA() IF ERROR:"
[1] "NNS.ARMA(... method = 'nonlin' , seasonal.factor = c( 17 ) ...)"
[1] "CURRENT nonlin OBJECTIVE FUNCTION = 6.43678662162006"
[1] "BEST method = 'nonlin' PATH MEMBER = c( 17 )"
[1] "BEST nonlin OBJECTIVE FUNCTION = 6.43678662162006"
[1] "CURRNET METHOD: both"
[1] "COPY LATEST PARAMETERS DIRECTLY FOR NNS.ARMA() IF ERROR:"
[1] "NNS.ARMA(... method = 'both' , seasonal.factor = c( 17 ) ...)"
[1] "CURRENT both OBJECTIVE FUNCTION = 5.80858375592161"
[1] "BEST method = 'both' PATH MEMBER = c( 17 )"
[1] "BEST both OBJECTIVE FUNCTION = 5.80858375592161"
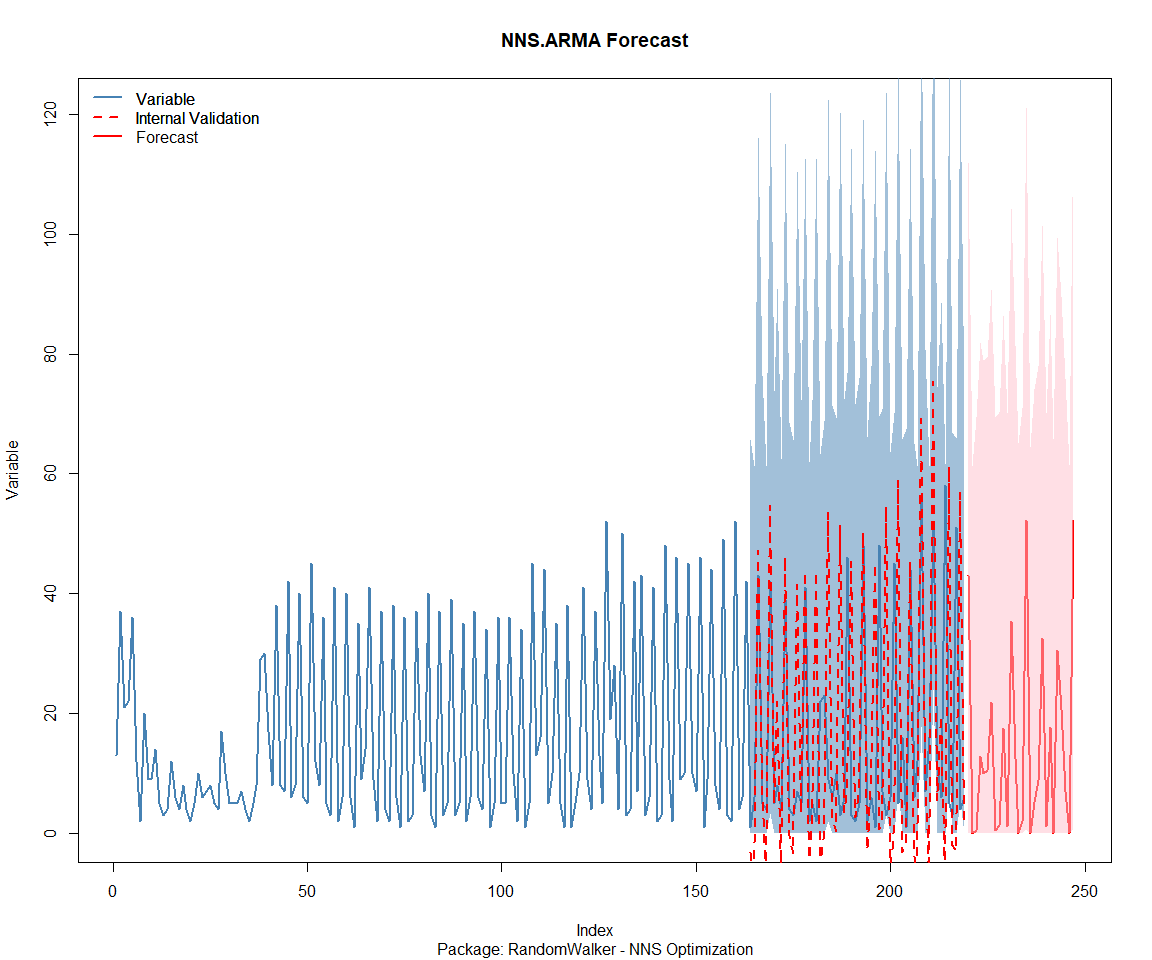
Package: tidyAML
[1] "CURRNET METHOD: lin"
[1] "COPY LATEST PARAMETERS DIRECTLY FOR NNS.ARMA() IF ERROR:"
[1] "NNS.ARMA(... method = 'lin' , seasonal.factor = c( 7 ) ...)"
[1] "CURRENT lin OBJECTIVE FUNCTION = 13.5878945558402"
[1] "BEST method = 'lin' PATH MEMBER = c( 7 )"
[1] "BEST lin OBJECTIVE FUNCTION = 13.5878945558402"
[1] "CURRNET METHOD: nonlin"
[1] "COPY LATEST PARAMETERS DIRECTLY FOR NNS.ARMA() IF ERROR:"
[1] "NNS.ARMA(... method = 'nonlin' , seasonal.factor = c( 7 ) ...)"
[1] "CURRENT nonlin OBJECTIVE FUNCTION = 6.28676155768198"
[1] "BEST method = 'nonlin' PATH MEMBER = c( 7 )"
[1] "BEST nonlin OBJECTIVE FUNCTION = 6.28676155768198"
[1] "CURRNET METHOD: both"
[1] "COPY LATEST PARAMETERS DIRECTLY FOR NNS.ARMA() IF ERROR:"
[1] "NNS.ARMA(... method = 'both' , seasonal.factor = c( 7 ) ...)"
[1] "CURRENT both OBJECTIVE FUNCTION = 6.78667422234949"
[1] "BEST method = 'both' PATH MEMBER = c( 7 )"
[1] "BEST both OBJECTIVE FUNCTION = 6.78667422234949"
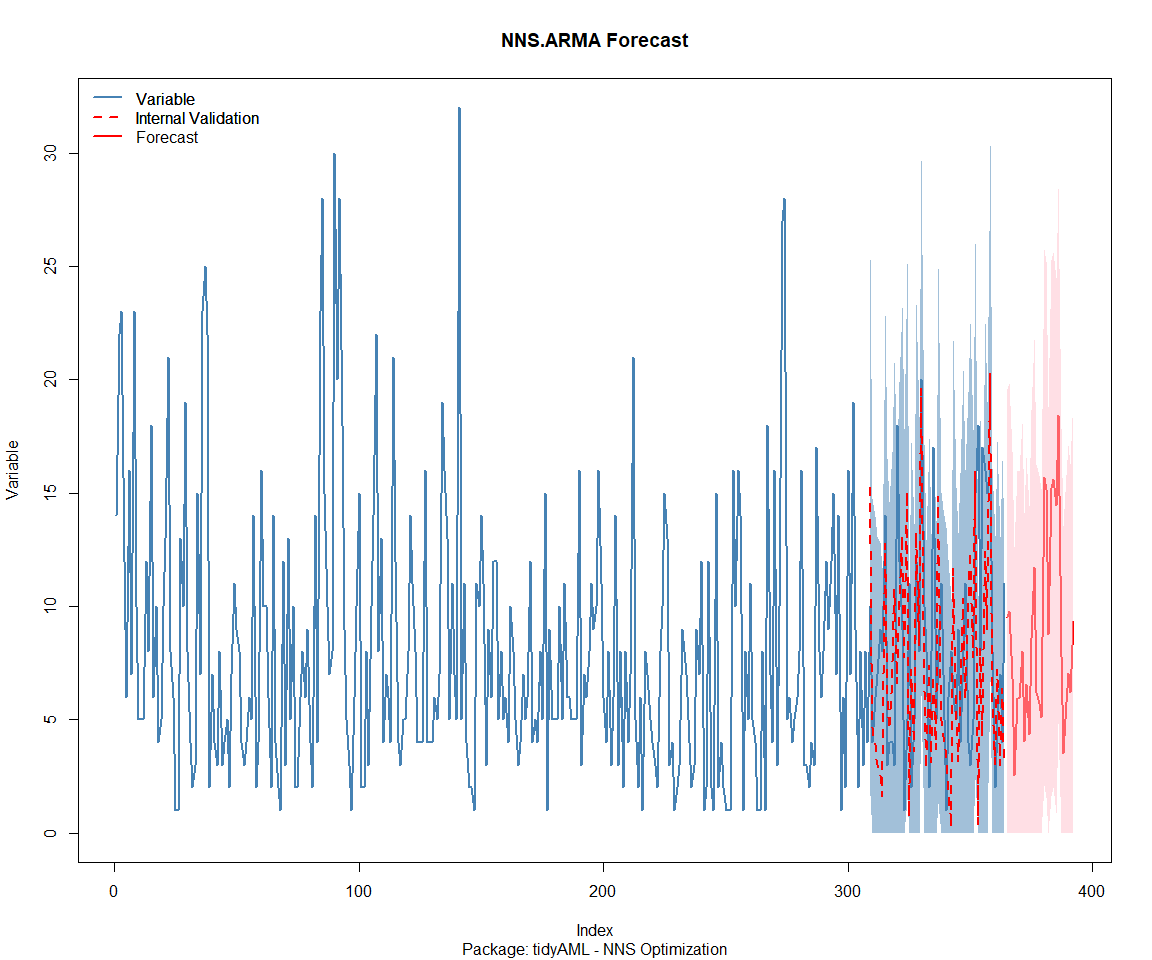
Package: TidyDensity
[1] "CURRNET METHOD: lin"
[1] "COPY LATEST PARAMETERS DIRECTLY FOR NNS.ARMA() IF ERROR:"
[1] "NNS.ARMA(... method = 'lin' , seasonal.factor = c( 17 ) ...)"
[1] "CURRENT lin OBJECTIVE FUNCTION = 6.09743366036836"
[1] "BEST method = 'lin' PATH MEMBER = c( 17 )"
[1] "BEST lin OBJECTIVE FUNCTION = 6.09743366036836"
[1] "CURRNET METHOD: nonlin"
[1] "COPY LATEST PARAMETERS DIRECTLY FOR NNS.ARMA() IF ERROR:"
[1] "NNS.ARMA(... method = 'nonlin' , seasonal.factor = c( 17 ) ...)"
[1] "CURRENT nonlin OBJECTIVE FUNCTION = 7.03871330774767"
[1] "BEST method = 'nonlin' PATH MEMBER = c( 17 )"
[1] "BEST nonlin OBJECTIVE FUNCTION = 7.03871330774767"
[1] "CURRNET METHOD: both"
[1] "COPY LATEST PARAMETERS DIRECTLY FOR NNS.ARMA() IF ERROR:"
[1] "NNS.ARMA(... method = 'both' , seasonal.factor = c( 17 ) ...)"
[1] "CURRENT both OBJECTIVE FUNCTION = 6.1196013268697"
[1] "BEST method = 'both' PATH MEMBER = c( 17 )"
[1] "BEST both OBJECTIVE FUNCTION = 6.1196013268697"
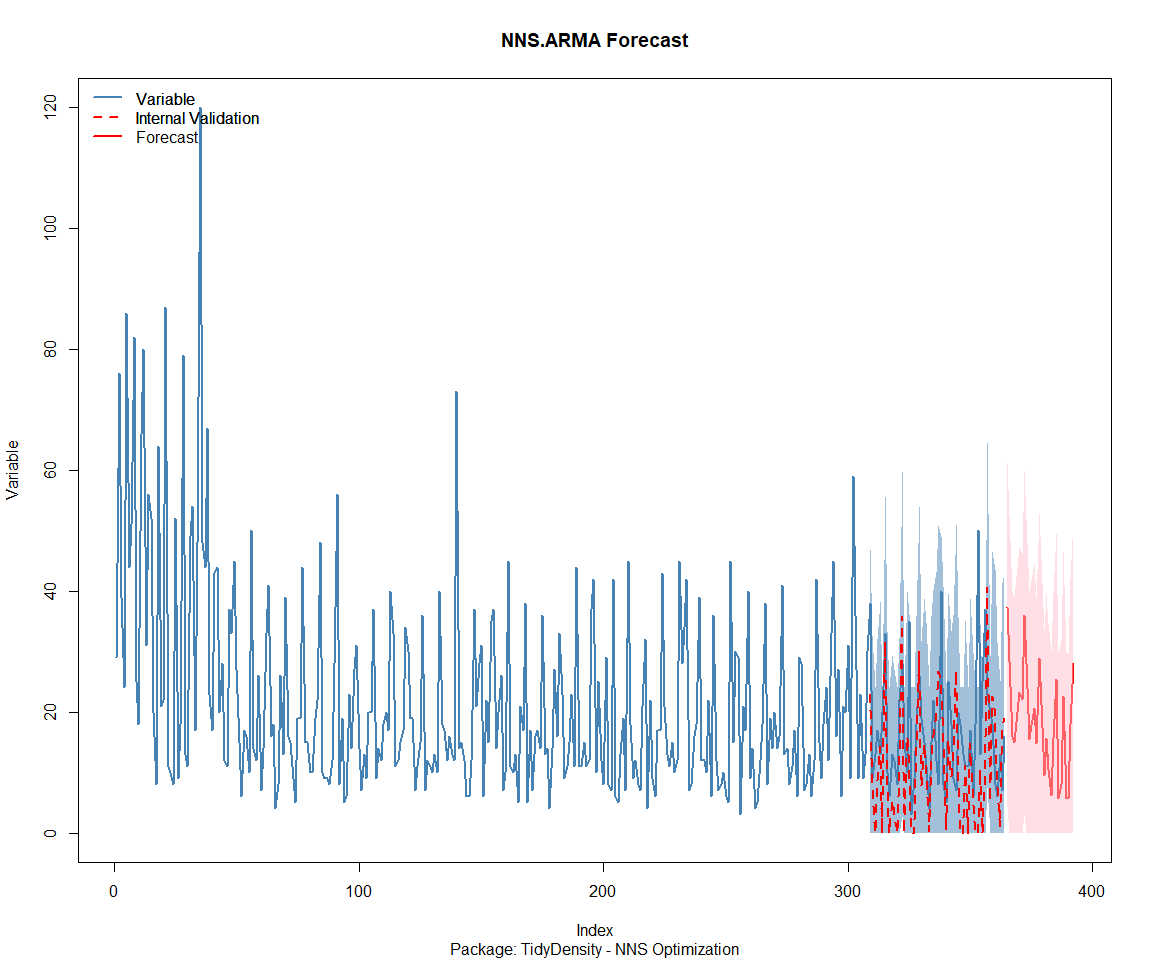
[[1]]
NULL
[[2]]
NULL
[[3]]
NULL
[[4]]
NULL
[[5]]
NULL
[[6]]
NULL
[[7]]
NULL
[[8]]
NULL
Pre-Processing
Now we are going to do some basic pre-processing.
data_padded_tbl <- base_data %>%
pad_by_time(
.date_var = date,
.pad_value = 0
)
# Get log interval and standardization parameters
log_params <- liv(data_padded_tbl$value, limit_lower = 0, offset = 1, silent = TRUE)
limit_lower <- log_params$limit_lower
limit_upper <- log_params$limit_upper
offset <- log_params$offset
data_liv_tbl <- data_padded_tbl %>%
# Get log interval transform
mutate(value_trans = liv(value, limit_lower = 0, offset = 1, silent = TRUE)$log_scaled)
# Get Standardization Params
std_params <- standard_vec(data_liv_tbl$value_trans, silent = TRUE)
std_mean <- std_params$mean
std_sd <- std_params$sd
data_transformed_tbl <- data_liv_tbl %>%
group_by(package) %>%
# get standardization
mutate(value_trans = standard_vec(value_trans, silent = TRUE)$standard_scaled) %>%
tk_augment_fourier(
.date_var = date,
.periods = c(7, 14, 30, 90, 180),
.K = 2
) %>%
tk_augment_timeseries_signature(
.date_var = date
) %>%
ungroup() %>%
select(-c(value, -year.iso))
Since this is panel data we can follow one of two different modeling strategies. We can search for a global model in the panel data or we can use nested forecasting finding the best model for each of the time series. Since we only have 5 panels, we will use nested forecasting.
To do this we will use the nest_timeseries and
split_nested_timeseries functions to create a nested tibble.
horizon <- 4*7
nested_data_tbl <- data_transformed_tbl %>%
# 0. Filter out column where package is NA
filter(!is.na(package)) %>%
# 1. Extending: We'll predict n days into the future.
extend_timeseries(
.id_var = package,
.date_var = date,
.length_future = horizon
) %>%
# 2. Nesting: We'll group by id, and create a future dataset
# that forecasts n days of extended data and
# an actual dataset that contains n*2 days
nest_timeseries(
.id_var = package,
.length_future = horizon
#.length_actual = horizon*2
) %>%
# 3. Splitting: We'll take the actual data and create splits
# for accuracy and confidence interval estimation of n das (test)
# and the rest is training data
split_nested_timeseries(
.length_test = horizon
)
nested_data_tbl
# A tibble: 8 × 4
package .actual_data .future_data .splits
<fct> <list> <list> <list>
1 healthyR.data <tibble [1,964 × 50]> <tibble [28 × 50]> <split [1936|28]>
2 healthyR <tibble [1,958 × 50]> <tibble [28 × 50]> <split [1930|28]>
3 healthyR.ts <tibble [1,894 × 50]> <tibble [28 × 50]> <split [1866|28]>
4 healthyverse <tibble [1,831 × 50]> <tibble [28 × 50]> <split [1803|28]>
5 healthyR.ai <tibble [1,700 × 50]> <tibble [28 × 50]> <split [1672|28]>
6 TidyDensity <tibble [1,551 × 50]> <tibble [28 × 50]> <split [1523|28]>
7 tidyAML <tibble [1,157 × 50]> <tibble [28 × 50]> <split [1129|28]>
8 RandomWalker <tibble [581 × 50]> <tibble [28 × 50]> <split [553|28]>
Now it is time to make some recipes and models using the modeltime workflow.
Modeltime Workflow
Recipe Object
recipe_base <- recipe(
value_trans ~ .
, data = extract_nested_test_split(nested_data_tbl)
)
recipe_base
recipe_date <- recipe(
value_trans ~ date
, data = extract_nested_test_split(nested_data_tbl)
)
Models
# Models ------------------------------------------------------------------
# Auto ARIMA --------------------------------------------------------------
model_spec_arima_no_boost <- arima_reg() %>%
set_engine(engine = "auto_arima")
wflw_auto_arima <- workflow() %>%
add_recipe(recipe = recipe_date) %>%
add_model(model_spec_arima_no_boost)
# NNETAR ------------------------------------------------------------------
model_spec_nnetar <- nnetar_reg(
mode = "regression"
, seasonal_period = "auto"
) %>%
set_engine("nnetar")
wflw_nnetar <- workflow() %>%
add_recipe(recipe = recipe_base) %>%
add_model(model_spec_nnetar)
# TSLM --------------------------------------------------------------------
model_spec_lm <- linear_reg() %>%
set_engine("lm")
wflw_lm <- workflow() %>%
add_recipe(recipe = recipe_base) %>%
add_model(model_spec_lm)
# MARS --------------------------------------------------------------------
model_spec_mars <- mars(mode = "regression") %>%
set_engine("earth")
wflw_mars <- workflow() %>%
add_recipe(recipe = recipe_date) %>%
add_model(model_spec_mars)
Nested Modeltime Tables
nested_modeltime_tbl <- modeltime_nested_fit(
# Nested Data
nested_data = nested_data_tbl,
control = control_nested_fit(
verbose = TRUE,
allow_par = FALSE
),
# Add workflows
wflw_auto_arima,
wflw_lm,
wflw_mars,
wflw_nnetar
)
nested_modeltime_tbl <- nested_modeltime_tbl[!is.na(nested_modeltime_tbl$package),]
Model Accuracy
nested_modeltime_tbl %>%
extract_nested_test_accuracy() %>%
filter(!is.na(package)) %>%
knitr::kable()
| package | .model_id | .model_desc | .type | mae | mape | mase | smape | rmse | rsq |
|---|---|---|---|---|---|---|---|---|---|
| healthyR.data | 1 | ARIMA | Test | 0.8159863 | 96.93802 | 0.7259502 | 159.58038 | 1.0496631 | 0.0005068 |
| healthyR.data | 2 | LM | Test | 0.9229448 | 124.53211 | 0.8211069 | 139.26910 | 1.2317849 | 0.0372048 |
| healthyR.data | 3 | EARTH | Test | 0.8382652 | 108.12613 | 0.7457709 | 148.46857 | 1.0720752 | 0.0113045 |
| healthyR.data | 4 | NNAR | Test | 0.8818935 | 110.82987 | 0.7845852 | 148.17993 | 1.1897878 | 0.0454629 |
| healthyR | 1 | ARIMA | Test | 0.7068904 | 1119.35918 | 0.7980669 | 134.08185 | 0.9707491 | 0.0498211 |
| healthyR | 2 | LM | Test | 0.8274818 | 979.87686 | 0.9342125 | 137.03718 | 1.1067681 | 0.0109756 |
| healthyR | 3 | EARTH | Test | 0.7288132 | 1054.76103 | 0.8228174 | 136.92960 | 0.9808697 | 0.0105180 |
| healthyR | 4 | NNAR | Test | 0.8493680 | 904.67335 | 0.9589216 | 156.30560 | 1.1372642 | 0.0606786 |
| healthyR.ts | 1 | ARIMA | Test | 0.6626943 | 416.28884 | 0.6549433 | 161.55928 | 0.8884590 | 0.0442957 |
| healthyR.ts | 2 | LM | Test | 0.7262949 | 466.97277 | 0.7178001 | 152.95209 | 0.9365532 | 0.0025874 |
| healthyR.ts | 3 | EARTH | Test | 0.6586476 | 462.58861 | 0.6509440 | 135.04014 | 0.8543500 | 0.0304667 |
| healthyR.ts | 4 | NNAR | Test | 0.7273269 | 306.86410 | 0.7188200 | 149.29508 | 0.9668539 | 0.0083291 |
| healthyverse | 1 | ARIMA | Test | 0.7036963 | 50.35908 | 1.1057562 | 55.81299 | 0.8036086 | 0.0009092 |
| healthyverse | 2 | LM | Test | 1.1787013 | 84.67273 | 1.8521575 | 127.36419 | 1.2927179 | 0.0455968 |
| healthyverse | 3 | EARTH | Test | 0.6335477 | 47.72804 | 0.9955279 | 48.99208 | 0.7454000 | 0.0245891 |
| healthyverse | 4 | NNAR | Test | 1.0768631 | 74.58750 | 1.6921336 | 114.20936 | 1.2029014 | 0.0402573 |
| healthyR.ai | 1 | ARIMA | Test | 0.5873011 | 95.16759 | 0.6958979 | 107.11901 | 0.8491322 | 0.0790989 |
| healthyR.ai | 2 | LM | Test | 0.7342010 | 165.37706 | 0.8699608 | 135.57288 | 0.9800540 | 0.0214529 |
| healthyR.ai | 3 | EARTH | Test | 0.7123549 | 127.57732 | 0.8440751 | 181.33310 | 0.8647833 | 0.0002226 |
| healthyR.ai | 4 | NNAR | Test | 0.6742027 | 138.39624 | 0.7988683 | 143.97035 | 0.8969802 | 0.0001942 |
| TidyDensity | 1 | ARIMA | Test | 1.1294611 | 119.16222 | 0.6666266 | 149.10254 | 1.3120093 | 0.0084879 |
| TidyDensity | 2 | LM | Test | 1.1857322 | 188.61427 | 0.6998387 | 149.68883 | 1.3125082 | 0.0242112 |
| TidyDensity | 3 | EARTH | Test | 1.2178074 | 178.59613 | 0.7187700 | 143.70424 | 1.3453572 | 0.0148620 |
| TidyDensity | 4 | NNAR | Test | 1.1359560 | 110.47944 | 0.6704600 | 159.66910 | 1.2564273 | 0.0570860 |
| tidyAML | 1 | ARIMA | Test | 0.7805752 | 187.88095 | 0.8674414 | 134.86361 | 1.1017829 | 0.0109321 |
| tidyAML | 2 | LM | Test | 0.7982927 | 328.33906 | 0.8871306 | 135.40246 | 1.0498433 | 0.0521735 |
| tidyAML | 3 | EARTH | Test | 2.7669464 | 1432.26251 | 3.0748656 | 157.25658 | 3.0645120 | 0.0299569 |
| tidyAML | 4 | NNAR | Test | 0.7423904 | 289.58870 | 0.8250073 | 126.41286 | 1.0545225 | 0.0513884 |
| RandomWalker | 1 | ARIMA | Test | 1.0165770 | 114.80505 | 0.7118413 | 159.97290 | 1.2180106 | 0.0164893 |
| RandomWalker | 2 | LM | Test | 0.9427253 | 102.24865 | 0.6601279 | 159.88847 | 1.0951189 | 0.0059683 |
| RandomWalker | 3 | EARTH | Test | 0.9500711 | 97.28127 | 0.6652716 | 178.12957 | 1.1240962 | 0.0339493 |
| RandomWalker | 4 | NNAR | Test | 0.9961610 | 114.57545 | 0.6975453 | 151.24628 | 1.1491339 | 0.0004538 |
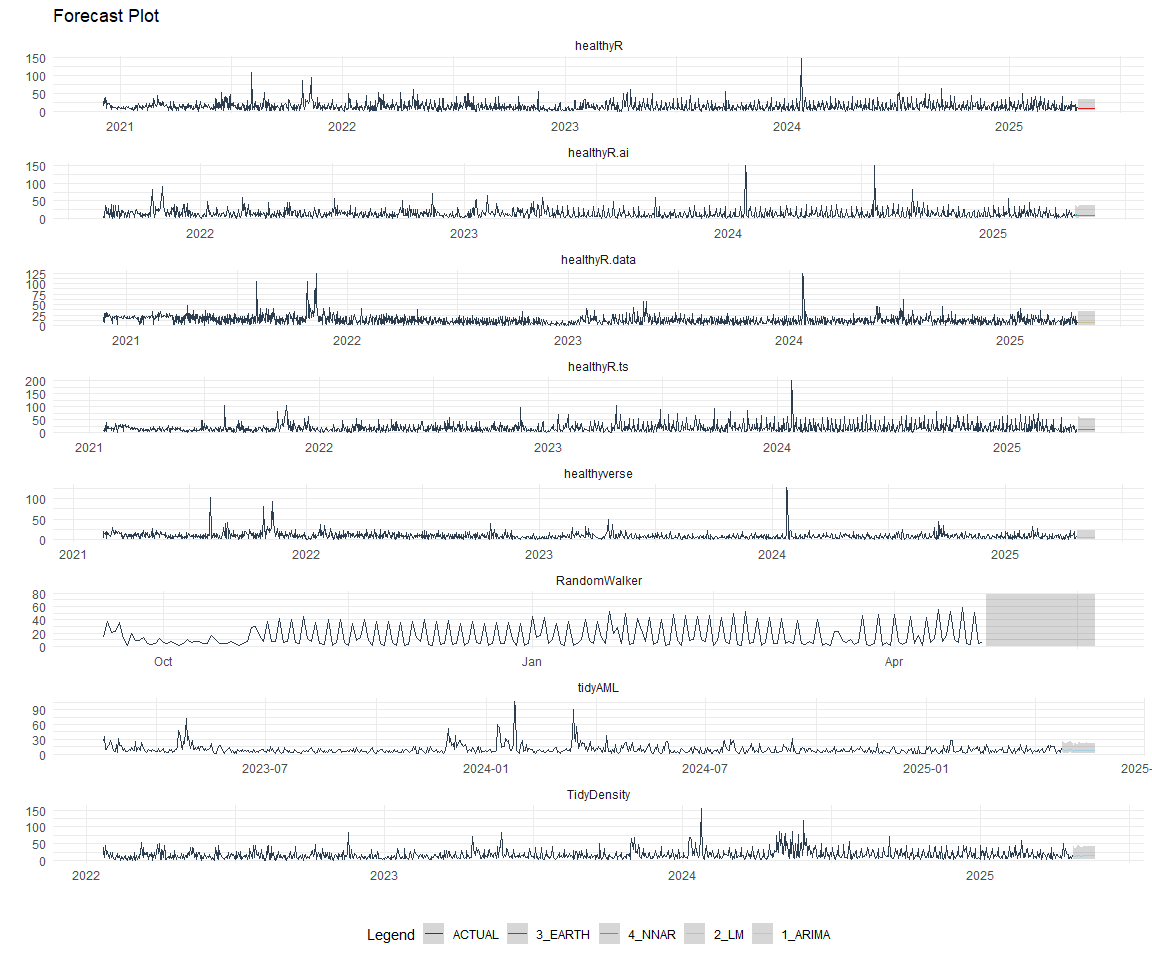
Plot Models
nested_modeltime_tbl %>%
extract_nested_test_forecast() %>%
group_by(package) %>%
filter_by_time(.date_var = .index, .start_date = max(.index) - 60) %>%
ungroup() %>%
plot_modeltime_forecast(
.interactive = FALSE,
.conf_interval_show = FALSE,
.facet_scales = "free"
) +
theme_minimal() +
facet_wrap(~ package, nrow = 3) +
theme(legend.position = "bottom")
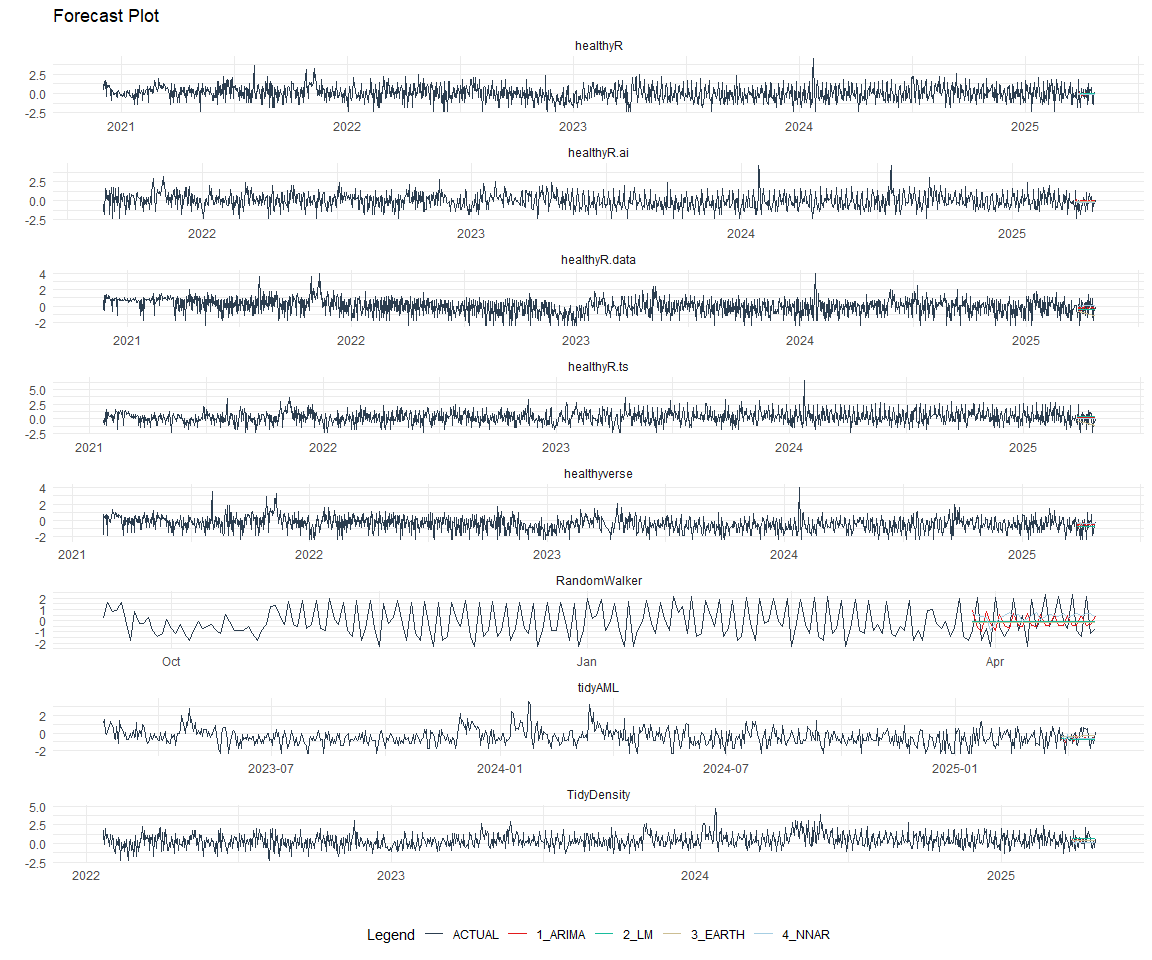
Best Model
best_nested_modeltime_tbl <- nested_modeltime_tbl %>%
modeltime_nested_select_best(
metric = "rmse",
minimize = TRUE,
filter_test_forecasts = TRUE
)
best_nested_modeltime_tbl %>%
extract_nested_best_model_report()
# Nested Modeltime Table
# A tibble: 8 × 10
package .model_id .model_desc .type mae mape mase smape rmse rsq
<fct> <int> <chr> <chr> <dbl> <dbl> <dbl> <dbl> <dbl> <dbl>
1 healthyR.d… 1 ARIMA Test 0.816 96.9 0.726 160. 1.05 5.07e-4
2 healthyR 1 ARIMA Test 0.707 1119. 0.798 134. 0.971 4.98e-2
3 healthyR.ts 3 EARTH Test 0.659 463. 0.651 135. 0.854 3.05e-2
4 healthyver… 3 EARTH Test 0.634 47.7 0.996 49.0 0.745 2.46e-2
5 healthyR.ai 1 ARIMA Test 0.587 95.2 0.696 107. 0.849 7.91e-2
6 TidyDensity 4 NNAR Test 1.14 110. 0.670 160. 1.26 5.71e-2
7 tidyAML 2 LM Test 0.798 328. 0.887 135. 1.05 5.22e-2
8 RandomWalk… 2 LM Test 0.943 102. 0.660 160. 1.10 5.97e-3
best_nested_modeltime_tbl %>%
extract_nested_test_forecast() %>%
#filter(!is.na(.model_id)) %>%
group_by(package) %>%
filter_by_time(.date_var = .index, .start_date = max(.index) - 60) %>%
ungroup() %>%
plot_modeltime_forecast(
.interactive = FALSE,
.conf_interval_alpha = 0.2,
.facet_scales = "free"
) +
facet_wrap(~ package, nrow = 3) +
theme_minimal() +
theme(legend.position = "bottom")
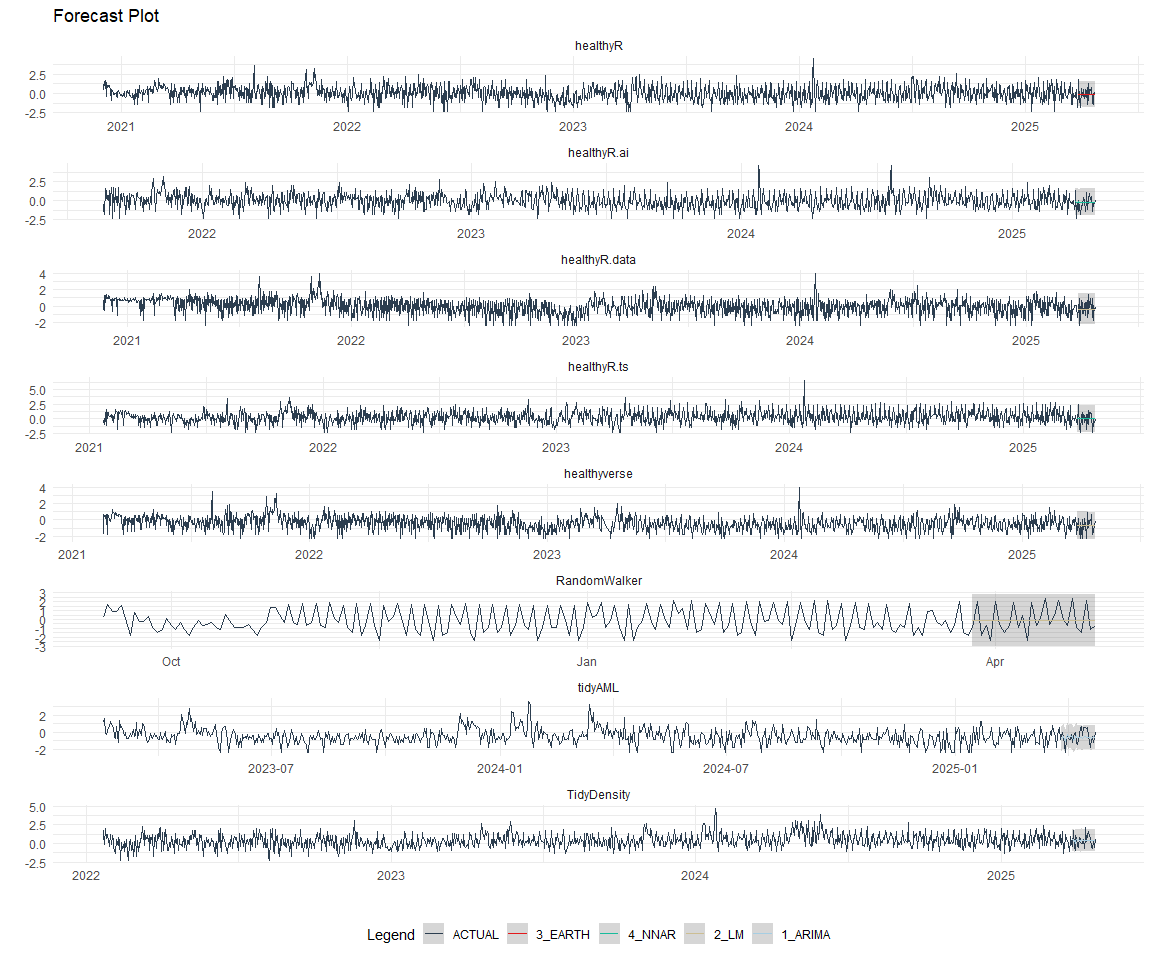
Refitting and Future Forecast
Now that we have the best models, we can make our future forecasts.
nested_modeltime_refit_tbl <- best_nested_modeltime_tbl %>%
modeltime_nested_refit(
control = control_nested_refit(verbose = TRUE)
)
nested_modeltime_refit_tbl
# Nested Modeltime Table
# A tibble: 8 × 5
package .actual_data .future_data .splits .modeltime_tables
<fct> <list> <list> <list> <list>
1 healthyR.data <tibble> <tibble> <split [1936|28]> <mdl_tm_t [1 × 5]>
2 healthyR <tibble> <tibble> <split [1930|28]> <mdl_tm_t [1 × 5]>
3 healthyR.ts <tibble> <tibble> <split [1866|28]> <mdl_tm_t [1 × 5]>
4 healthyverse <tibble> <tibble> <split [1803|28]> <mdl_tm_t [1 × 5]>
5 healthyR.ai <tibble> <tibble> <split [1672|28]> <mdl_tm_t [1 × 5]>
6 TidyDensity <tibble> <tibble> <split [1523|28]> <mdl_tm_t [1 × 5]>
7 tidyAML <tibble> <tibble> <split [1129|28]> <mdl_tm_t [1 × 5]>
8 RandomWalker <tibble> <tibble> <split [553|28]> <mdl_tm_t [1 × 5]>
nested_modeltime_refit_tbl %>%
extract_nested_future_forecast() %>%
group_by(package) %>%
mutate(across(.value:.conf_hi, .fns = ~ standard_inv_vec(
x = .,
mean = std_mean,
sd = std_sd
)$standard_inverse_value)) %>%
mutate(across(.value:.conf_hi, .fns = ~ liiv(
x = .,
limit_lower = limit_lower,
limit_upper = limit_upper,
offset = offset
)$rescaled_v)) %>%
filter_by_time(.date_var = .index, .start_date = max(.index) - 60) %>%
ungroup() %>%
plot_modeltime_forecast(
.interactive = FALSE,
.conf_interval_alpha = 0.2,
.facet_scales = "free"
) +
facet_wrap(~ package, nrow = 3) +
theme_minimal() +
theme(legend.position = "bottom")